+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7jjf | ||||||
|---|---|---|---|---|---|---|---|
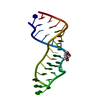
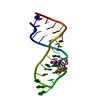
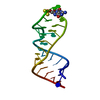
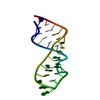
| Title | Sarcin-ricin loop with modified residue. | ||||||
 Components Components | RNA/DNA (27-mer) | ||||||
 Keywords Keywords | DNA-RNA HYBRID / sarcin-ricin loop / RNA crystal structure / RNA tetraloops / RNA | ||||||
| Function / homology | RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.2 Å MOLECULAR REPLACEMENT / Resolution: 1.2 Å | ||||||
 Authors Authors | Harp, J.M. / Pallan, P.S. / Egli, M. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2020 Journal: Biochemistry / Year: 2020Title: Incorporating a Thiophosphate Modification into a Common RNA Tetraloop Motif Causes an Unanticipated Stability Boost. Authors: Pallan, P.S. / Lybrand, T.P. / Schlegel, M.K. / Harp, J.M. / Jahns, H. / Manoharan, M. / Egli, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7jjf.cif.gz 7jjf.cif.gz | 51.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7jjf.ent.gz pdb7jjf.ent.gz | 36.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7jjf.json.gz 7jjf.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jj/7jjf https://data.pdbj.org/pub/pdb/validation_reports/jj/7jjf ftp://data.pdbj.org/pub/pdb/validation_reports/jj/7jjf ftp://data.pdbj.org/pub/pdb/validation_reports/jj/7jjf | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7jjdC  7jjeC  3dvzS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 8726.282 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  |
|---|---|
| #2: Chemical | ChemComp-MG / |
| #3: Water | ChemComp-HOH / |
| Has ligand of interest | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.81 Å3/Da / Density % sol: 32.06 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop / pH: 7 Details: RNA was dissolved in 1 mM Na2 EDTA (pH 8.0) and 10 mM Tris-HCl (pH 8.0) to attain 350 uM final RNA concentration. The RNA sample was annealed by heating at 65 C for 2 min and slowly cooling ...Details: RNA was dissolved in 1 mM Na2 EDTA (pH 8.0) and 10 mM Tris-HCl (pH 8.0) to attain 350 uM final RNA concentration. The RNA sample was annealed by heating at 65 C for 2 min and slowly cooling to room temperature. Crystallization set-ups were made by mixing 4 uL of RNA solution with 2 uL of a crystallization buffer composed of 3.0 M ammonium sulfate, 10 mM magnesium chloride, 10 mM manganese chloride, and 50 mM potassium 3-(N-morpholino) propanesulfonic acid (MOPS), pH 7.0 at 18 C. Crystals appear in about two weeks time. |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source: LIQUID ANODE / Type: Excillum MetalJet D2+ 70 kV / Wavelength: 1.3418 Å |
| Detector | Type: Bruker PHOTON III / Detector: PIXEL / Date: Mar 19, 2020 |
| Radiation | Monochromator: multi-layer confocal / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.3418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.2→27.36 Å / % possible obs: 94.1 % / Redundancy: 17 % / Biso Wilson estimate: 4.79 Å2 / Rmerge(I) obs: 0.089 / Net I/σ(I): 23.47 |
| Reflection shell | Resolution: 1.2→1.24 Å |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3DVZ Resolution: 1.2→27.36 Å / Cross valid method: FREE R-VALUE / σ(F): 4.58 / Phase error: 41.508 Stereochemistry target values: GEOSTD + MONOMER LIBRARY + CDL V1.2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 10.76 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.2→27.36 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller







 PDBj
PDBj
































