+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1cm1 | ||||||
|---|---|---|---|---|---|---|---|
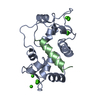
| Title | MOTIONS OF CALMODULIN-SINGLE-CONFORMER REFINEMENT | ||||||
 Components Components |
| ||||||
 Keywords Keywords | COMPLEX (CALCIUM-BINDING/TRANSFERASE) / COMPLEX (CALCIUM-BINDING-TRANSFERASE) / EF-HAND CALCIUM-BINDING PROTEIN / COMPLEX (CALCIUM-BINDING-TRANSFERASE) complex | ||||||
| Function / homology |  Function and homology information Function and homology informationHSF1-dependent transactivation / regulation of synaptic vesicle docking / glutamatergic postsynaptic density / RAF activation / Ion transport by P-type ATPases /  regulation of store-operated calcium channel activity / peptidyl-threonine autophosphorylation / regulation of endocannabinoid signaling pathway / calcium- and calmodulin-dependent protein kinase complex / regulation of high voltage-gated calcium channel activity ...HSF1-dependent transactivation / regulation of synaptic vesicle docking / glutamatergic postsynaptic density / RAF activation / Ion transport by P-type ATPases / regulation of store-operated calcium channel activity / peptidyl-threonine autophosphorylation / regulation of endocannabinoid signaling pathway / calcium- and calmodulin-dependent protein kinase complex / regulation of high voltage-gated calcium channel activity ...HSF1-dependent transactivation / regulation of synaptic vesicle docking / glutamatergic postsynaptic density / RAF activation / Ion transport by P-type ATPases /  regulation of store-operated calcium channel activity / peptidyl-threonine autophosphorylation / regulation of endocannabinoid signaling pathway / calcium- and calmodulin-dependent protein kinase complex / regulation of high voltage-gated calcium channel activity / calmodulin dependent kinase signaling pathway / Interferon gamma signaling / calcium-dependent protein serine/threonine kinase activity / NMDA selective glutamate receptor signaling pathway / regulation of store-operated calcium channel activity / peptidyl-threonine autophosphorylation / regulation of endocannabinoid signaling pathway / calcium- and calmodulin-dependent protein kinase complex / regulation of high voltage-gated calcium channel activity / calmodulin dependent kinase signaling pathway / Interferon gamma signaling / calcium-dependent protein serine/threonine kinase activity / NMDA selective glutamate receptor signaling pathway /  regulation of neuron migration / regulation of neuron migration /  : / : /  Ca2+/calmodulin-dependent protein kinase / regulation of response to tumor cell / positive regulation of autophagic cell death / DAPK1-calmodulin complex / regulation of neurotransmitter secretion / dendritic spine development / : / establishment of protein localization to mitochondrial membrane / Trafficking of AMPA receptors / type 3 metabotropic glutamate receptor binding / Ca2+ pathway / positive regulation of calcium ion transport / postsynaptic neurotransmitter receptor diffusion trapping / postsynaptic specialization membrane / presynaptic cytosol / negative regulation of hydrolase activity / establishment of protein localization to membrane / calmodulin-dependent protein kinase activity / RAF/MAP kinase cascade / GTPase activating protein binding / dendrite morphogenesis / regulation of mitochondrial membrane permeability involved in apoptotic process / regulation of neurotransmitter receptor localization to postsynaptic specialization membrane / postsynaptic cytosol / regulation of synaptic vesicle endocytosis / negative regulation of high voltage-gated calcium channel activity / regulation of synaptic vesicle exocytosis / negative regulation of calcium ion export across plasma membrane / organelle localization by membrane tethering / regulation of neuronal synaptic plasticity / regulation of cardiac muscle cell action potential / mitochondrion-endoplasmic reticulum membrane tethering / Ion homeostasis / autophagosome membrane docking / positive regulation of ryanodine-sensitive calcium-release channel activity / Ca2+/calmodulin-dependent protein kinase / regulation of response to tumor cell / positive regulation of autophagic cell death / DAPK1-calmodulin complex / regulation of neurotransmitter secretion / dendritic spine development / : / establishment of protein localization to mitochondrial membrane / Trafficking of AMPA receptors / type 3 metabotropic glutamate receptor binding / Ca2+ pathway / positive regulation of calcium ion transport / postsynaptic neurotransmitter receptor diffusion trapping / postsynaptic specialization membrane / presynaptic cytosol / negative regulation of hydrolase activity / establishment of protein localization to membrane / calmodulin-dependent protein kinase activity / RAF/MAP kinase cascade / GTPase activating protein binding / dendrite morphogenesis / regulation of mitochondrial membrane permeability involved in apoptotic process / regulation of neurotransmitter receptor localization to postsynaptic specialization membrane / postsynaptic cytosol / regulation of synaptic vesicle endocytosis / negative regulation of high voltage-gated calcium channel activity / regulation of synaptic vesicle exocytosis / negative regulation of calcium ion export across plasma membrane / organelle localization by membrane tethering / regulation of neuronal synaptic plasticity / regulation of cardiac muscle cell action potential / mitochondrion-endoplasmic reticulum membrane tethering / Ion homeostasis / autophagosome membrane docking / positive regulation of ryanodine-sensitive calcium-release channel activity /  nitric-oxide synthase binding / positive regulation of cardiac muscle cell apoptotic process / protein phosphatase activator activity / calcium channel regulator activity / positive regulation of cyclic-nucleotide phosphodiesterase activity / positive regulation of phosphoprotein phosphatase activity / nitric-oxide synthase binding / positive regulation of cardiac muscle cell apoptotic process / protein phosphatase activator activity / calcium channel regulator activity / positive regulation of cyclic-nucleotide phosphodiesterase activity / positive regulation of phosphoprotein phosphatase activity /  adenylate cyclase binding / adenylate cyclase binding /  catalytic complex / detection of calcium ion / negative regulation of ryanodine-sensitive calcium-release channel activity / Unblocking of NMDA receptors, glutamate binding and activation / catalytic complex / detection of calcium ion / negative regulation of ryanodine-sensitive calcium-release channel activity / Unblocking of NMDA receptors, glutamate binding and activation /  glutamate receptor binding / regulation of cardiac muscle contraction / calcium channel inhibitor activity / cellular response to interferon-beta / positive regulation of DNA binding / enzyme regulator activity / regulation of protein localization to plasma membrane / glutamate receptor binding / regulation of cardiac muscle contraction / calcium channel inhibitor activity / cellular response to interferon-beta / positive regulation of DNA binding / enzyme regulator activity / regulation of protein localization to plasma membrane /  voltage-gated potassium channel complex / voltage-gated potassium channel complex /  phosphatidylinositol 3-kinase binding / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / regulation of calcium-mediated signaling / positive regulation of protein dephosphorylation / phosphatidylinositol 3-kinase binding / regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / regulation of calcium-mediated signaling / positive regulation of protein dephosphorylation /  titin binding / regulation of ryanodine-sensitive calcium-release channel activity / potassium ion transmembrane transport / response to amphetamine / titin binding / regulation of ryanodine-sensitive calcium-release channel activity / potassium ion transmembrane transport / response to amphetamine /  calcium channel complex / activation of adenylate cyclase activity / adenylate cyclase activator activity / ionotropic glutamate receptor signaling pathway / dendrite cytoplasm / calcium channel complex / activation of adenylate cyclase activity / adenylate cyclase activator activity / ionotropic glutamate receptor signaling pathway / dendrite cytoplasm /  regulation of heart rate / nitric-oxide synthase regulator activity / regulation of heart rate / nitric-oxide synthase regulator activity /  sarcomere / sarcomere /  regulation of cytokinesis / positive regulation of nitric-oxide synthase activity / response to ischemia / calcium-mediated signaling / spindle microtubule / angiotensin-activated signaling pathway / G1/S transition of mitotic cell cycle / positive regulation of receptor signaling pathway via JAK-STAT / Schaffer collateral - CA1 synapse / regulation of cytokinesis / positive regulation of nitric-oxide synthase activity / response to ischemia / calcium-mediated signaling / spindle microtubule / angiotensin-activated signaling pathway / G1/S transition of mitotic cell cycle / positive regulation of receptor signaling pathway via JAK-STAT / Schaffer collateral - CA1 synapse /  spindle pole / cellular response to type II interferon / response to calcium ion / calcium-dependent protein binding / calcium ion transport spindle pole / cellular response to type II interferon / response to calcium ion / calcium-dependent protein binding / calcium ion transportSimilarity search - Function | ||||||
| Biological species |   Bos taurus (cattle) Bos taurus (cattle) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Wall, M.E. / Phillips Jr., G.N. | ||||||
 Citation Citation |  Journal: Structure / Year: 1997 Journal: Structure / Year: 1997Title: Motions of calmodulin characterized using both Bragg and diffuse X-ray scattering. Authors: Wall, M.E. / Clarage, J.B. / Phillips Jr., G.N. #1:  Journal: Science / Year: 1993 Journal: Science / Year: 1993Title: Modulation of Calmodulin Plasticity in Molecular Recognition on the Basis of X-Ray Structures Authors: Meador, W.E. / Means, A.R. / Quiocho, F.A. #2:  Journal: Science / Year: 1992 Journal: Science / Year: 1992Title: Target Enzyme Recognition by Calmodulin: 2.4 A Structure of a Calmodulin-Peptide Complex Authors: Meador, W.E. / Means, A.R. / Quiocho, F.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1cm1.cif.gz 1cm1.cif.gz | 44 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1cm1.ent.gz pdb1cm1.ent.gz | 32.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1cm1.json.gz 1cm1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/cm/1cm1 https://data.pdbj.org/pub/pdb/validation_reports/cm/1cm1 ftp://data.pdbj.org/pub/pdb/validation_reports/cm/1cm1 ftp://data.pdbj.org/pub/pdb/validation_reports/cm/1cm1 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 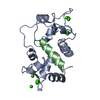 1cm4C  1cdlS  1cdmS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein |  Mass: 16721.350 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Bos taurus (cattle) / Description: SIGMA LOT 54H9558 / Organ: BRAIN Bos taurus (cattle) / Description: SIGMA LOT 54H9558 / Organ: BRAIN / References: UniProt: P02593, UniProt: P62157*PLUS / References: UniProt: P02593, UniProt: P62157*PLUS | ||
|---|---|---|---|
| #2: Protein/peptide | Mass: 2886.528 Da / Num. of mol.: 1 / Fragment: CALMODULIN BINDING DOMAIN, RESIDUES 290 - 314 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Bos taurus (cattle) / References: UniProt: P11275, EC: 2.7.1.123 Bos taurus (cattle) / References: UniProt: P11275, EC: 2.7.1.123 | ||
| #3: Chemical | ChemComp-CA / #4: Water | ChemComp-HOH / |  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.23 Å3/Da / Density % sol: 44.86 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | Method: vapor diffusion - hanging drop - microseeding / pH: 5.2 Details: DIFFRACTION-QUALITY CRYSTALS WERE MICROSEEDED IN HANGING DROPS OVER 100 MM SODIUM ACETATE AT PH 5.2, WITH 20% POLY-ETHYLENE GLYCOL 6000 (PEG 6000), 10 MM CALCIUM CHLORIDE AND 0.02% SODIUM ...Details: DIFFRACTION-QUALITY CRYSTALS WERE MICROSEEDED IN HANGING DROPS OVER 100 MM SODIUM ACETATE AT PH 5.2, WITH 20% POLY-ETHYLENE GLYCOL 6000 (PEG 6000), 10 MM CALCIUM CHLORIDE AND 0.02% SODIUM AZIDE. STOCK SOLUTIONS OF 24 MG/ML BOVINE BRAIN CALMODULIN (SIGMA LOT 54H9558), 14 MG/ML CAMKII-ALPHA PEPTIDE, AND 30% PEG WERE MIXED INTO HANGING DROPS IN ABOUT A 4-2-1 RATIO., vapor diffusion - hanging drop - microseeding | ||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop / Details: used to seeding | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 302 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CHESS CHESS  / Beamline: F2 / Wavelength: 0.98 / Beamline: F2 / Wavelength: 0.98 |
| Detector | Type: PRINCETON 2K / Detector: CCD / Date: May 1, 1996 / Details: MIRRORS |
| Radiation | Monochromator: SINGLE-CRYSTAL / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.98 Å / Relative weight: 1 : 0.98 Å / Relative weight: 1 |
| Reflection | Resolution: 2→10 Å / Num. obs: 11522 / % possible obs: 92.1 % / Observed criterion σ(I): 2 / Biso Wilson estimate: 22.5 Å2 / Rsym value: 0.061 / Net I/σ(I): 9.2 |
| Reflection shell | Resolution: 2→2.07 Å / Rsym value: 0.24 / % possible all: 82.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRIES 1CDM AND 1CDL Resolution: 2→10 Å / Rfactor Rfree error: 0.009 / Data cutoff high absF: 100000 / Data cutoff low absF: 0.1 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 2 Details: RESIDUES 74 - 83, WHICH ARE NOT PRESENT IN PDB ENTRY 1CDM, WERE ADDED FOR THIS REFINEMENT. AN INITIAL GUESS WAS OBTAINED FROM THE COORDINATES OF PDB ENTRY 1CDL. THESE RESIDUES DO NOT SHOW ...Details: RESIDUES 74 - 83, WHICH ARE NOT PRESENT IN PDB ENTRY 1CDM, WERE ADDED FOR THIS REFINEMENT. AN INITIAL GUESS WAS OBTAINED FROM THE COORDINATES OF PDB ENTRY 1CDL. THESE RESIDUES DO NOT SHOW CONNECTED ELECTRON DENSITY AT A LEVEL OF 1SIGMA.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 41.4 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.07 Å / Rfactor Rfree error: 0.033 / Total num. of bins used: 10
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.851 / Classification: refinement X-PLOR / Version: 3.851 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj








