+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDCP3 |
|---|---|
 Sample Sample | Proline utilization A from Bdellovibrio bacteriovorus
|
| Function / homology |  Function and homology information Function and homology informationproline dehydrogenase activity / L-glutamate gamma-semialdehyde dehydrogenase / L-glutamate gamma-semialdehyde dehydrogenase activity / : / cytoplasmic side of plasma membrane / DNA-binding transcription factor activity Similarity search - Function |
| Biological species |  Bdellovibrio bacteriovorus (strain ATCC 15356 / DSM 50701 / NCIB 9529 / HD100) (bacteria) Bdellovibrio bacteriovorus (strain ATCC 15356 / DSM 50701 / NCIB 9529 / HD100) (bacteria) |
 Citation Citation |  Journal: FEBS J / Year: 2017 Journal: FEBS J / Year: 2017Title: Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure. Authors: David A Korasick / Harkewal Singh / Travis A Pemberton / Min Luo / Richa Dhatwalia / John J Tanner /  Abstract: Many enzymes form homooligomers, yet the functional significance of self-association is seldom obvious. Herein, we examine the connection between oligomerization and catalytic function for proline ...Many enzymes form homooligomers, yet the functional significance of self-association is seldom obvious. Herein, we examine the connection between oligomerization and catalytic function for proline utilization A (PutA) enzymes. PutAs are bifunctional enzymes that catalyze both reactions of proline catabolism. Type A PutAs are the smallest members of the family, possessing a minimal domain architecture consisting of N-terminal proline dehydrogenase and C-terminal l-glutamate-γ-semialdehyde dehydrogenase modules. Type A PutAs form domain-swapped dimers, and in one case (Bradyrhizobium japonicum PutA), two of the dimers assemble into a ring-shaped tetramer. Whereas the dimer has a clear role in substrate channeling, the functional significance of the tetramer is unknown. To address this question, we performed structural studies of four-type A PutAs from two clades of the PutA tree. The crystal structure of Bdellovibrio bacteriovorus PutA covalently inactivated by N-propargylglycine revealed a fold and substrate-channeling tunnel similar to other PutAs. Small-angle X-ray scattering (SAXS) and analytical ultracentrifugation indicated that Bdellovibrio PutA is dimeric in solution, in contrast to the prediction from crystal packing of a stable tetrameric assembly. SAXS studies of two other type A PutAs from separate clades also suggested that the dimer predominates in solution. To assess whether the tetramer of B. japonicum PutA is necessary for catalytic function, a hot spot disruption mutant that cleanly produces dimeric protein was generated. The dimeric variant exhibited kinetic parameters similar to the wild-type enzyme. These results implicate the domain-swapped dimer as the core structural and functional unit of type A PutAs. ENZYMES: Proline dehydrogenase (EC 1.5.5.2); l-glutamate-γ-semialdehyde dehydrogenase (EC 1.2.1.88). DATABASES: The atomic coordinates and structure factor amplitudes have been deposited in the Protein Data Bank under accession number 5UR2. The SAXS data have been deposited in the SASBDB under the ...DATABASES: The atomic coordinates and structure factor amplitudes have been deposited in the Protein Data Bank under accession number 5UR2. The SAXS data have been deposited in the SASBDB under the following accession codes: SASDCP3 (BbPutA), SASDCQ3 (DvPutA 1.5 mg·mL ), SASDCX3 (DvPutA 3.0 mg·mL ), SASDCY3 (DvPutA 4.5 mg·mL ), SASDCR3 (LpPutA 3.0 mg·mL ), SASDCV3 (LpPutA 5.0 mg·mL ), SASDCW3 (LpPutA 8.0 mg·mL ), SASDCS3 (BjPutA 2.3 mg·mL ), SASDCT3 (BjPutA 4.7 mg·mL ), SASDCU3 (BjPutA 7.0 mg·mL ), SASDCZ3 (R51E 2.3 mg·mL ), SASDC24 (R51E 4.7 mg·mL ), SASDC34 (R51E 7.0 mg·mL ). |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDCP3 SASDCP3 |
|---|
-Related structure data
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
| Model #1419 |  Type: dummy / Radius of dummy atoms: 2.90 A / Symmetry: P2 / Chi-square value: 1.168  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|---|
| Model #1420 |  Type: dummy / Software: (5.0 r6293) / Radius of dummy atoms: 4.25 A / Chi-square value: 1.168  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
- Sample
Sample
 Sample Sample | Name: Proline utilization A from Bdellovibrio bacteriovorus |
|---|---|
| Buffer | Name: 50 mM Tris, 125 mM NaCl, 1 mM EDTA, and 1 mM tris(3-hydroxypropyl)phosphine (THP) at pH 7.5, pH: 7.5 |
| Entity #678 | Type: protein / Description: Bifunctional protein PutA / Formula weight: 109.504 / Num. of mol.: 2 Source: Bdellovibrio bacteriovorus (strain ATCC 15356 / DSM 50701 / NCIB 9529 / HD100) References: UniProt: Q6MNK1 Sequence: GHMNDIQSQI VSRGEEILKR MESQSKASIF SKDFWYGSIM EWSMKNEKFK TNMFRFVDVL PSINSGDEVA RHLKEYFSED GGTLPPVFNV GLGLGSLAPG LMAGAIKKNV MGMAKMFITG ESPDEALPVL KKARKNKMTF TVDILGEATL SEKEAQDYSN KYMELVTWLA ...Sequence: GHMNDIQSQI VSRGEEILKR MESQSKASIF SKDFWYGSIM EWSMKNEKFK TNMFRFVDVL PSINSGDEVA RHLKEYFSED GGTLPPVFNV GLGLGSLAPG LMAGAIKKNV MGMAKMFITG ESPDEALPVL KKARKNKMTF TVDILGEATL SEKEAQDYSN KYMELVTWLA KDAEKWDEVP QIDRDHEGAL PKVNVSVKMT ALYSQIKDAA WDESKKILKD RLRPVFRLGM EKGVFVNLDM EQYSVKHLTL EVFTELINEP EFKNYKFFGI VIQAYLRDSF EDVKSLTEFA QKRGTPFWVR LVKGAYWDYE TIEAEQRGWP VPVYTNKAES DANYELCAKY LLENIKFIRP AFASHNVRTL AACMLYAEKL NIPKEALEFQ MLYGMAEPIK KTIVDMGYRM REYAPVGELI PGMAYLVRRL LENTSNESWL RGKFADNKSM AELLKDPAQG LTPTSPVIPK KPGKFYNEPL LDFAVKADRE KMLKALAEAK ASLPVNVNIV INNKELQSGK IFDRVNPSQS DQIVGKIQMA TTEQAEQAMQ AAQTAYKTWK NVPCEQRAAL VDKLADIMTR DRFKLIATQV LEVGKPWAEA DGDIGEAIDF CRYYARHMRE LQKPLRVGGL PGELSHYIYK SRGVTAVIAP WNFPLAILAG MVTAAAVAGN TVVMKPAEQS TVVAWGLMKM IQEAGFPQGV INFLPGYGEE VGEYIVNHKY TTTIAFTGSK AVGLHIMNRA AVVQPGQQHV KRCIIEMGGK NAVIIDNDAD LDEAVDGVIY SAFGFSGQKC SAASRVIVLD EVYDRFVDRL VETAKSIEIH PAENPKAYMG PVVDKEAYDR ILGTIAEAEK NHKLLFKGSV PGGGFFAPPT IFGDVPGDAK LAQAEIFGPV VAVIRAKNLD QALDIANSTE YALTGGVFSR SPANINRVKE ELEVGNLYVN RGITGAMVDR HPFGGFKMSG IGSKTGGPDY LKQYMEPACV TENTLRRGFA PAEE |
-Experimental information
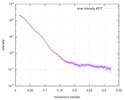
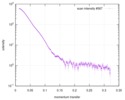
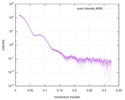
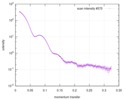
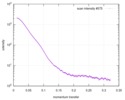
| Beam | Instrument name: Advanced Light Source (ALS) 12.3.1 (SIBYLS) City: Berkeley, CA / 国: USA  / Type of source: X-ray synchrotron / Type of source: X-ray synchrotron | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: MAR 165 CCD | |||||||||||||||||||||||||||||||||
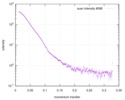
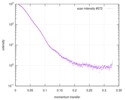
| Scan | Measurement date: Jun 8, 2012 / Unit: 1/A /
| |||||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| |||||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller