[English] 日本語
 Yorodumi
Yorodumi- PDB-7lj0: Linker 3 and scaffolded phycoerythrin beta subunit from the phyco... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7lj0 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
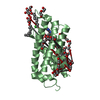
| Title | Linker 3 and scaffolded phycoerythrin beta subunit from the phycobilisome of Porphyridium purpureum | ||||||||||||
 Components Components |
| ||||||||||||
 Keywords Keywords | PHOTOSYNTHESIS / linker / phycobilisome / PBS / light harvesting / red algae / scaffolding | ||||||||||||
| Function / homology |  Function and homology information Function and homology information | ||||||||||||
| Biological species |  Porphyridium purpureum (eukaryote) Porphyridium purpureum (eukaryote) | ||||||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 2.8 Å | ||||||||||||
 Authors Authors | Rathbone, H.W. / Landsberg, M.J. / Michie, K.A. / Green, B.R. / Curmi, P.M.G. | ||||||||||||
| Funding support |  Australia, Australia,  United States, 3items United States, 3items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structural basis of energy transfer in Porphyridium purpureum phycobilisome. Authors: Jianfei Ma / Xin You / Shan Sun / Xiaoxiao Wang / Song Qin / Sen-Fang Sui /  Abstract: Photosynthetic organisms have developed various light-harvesting systems to adapt to their environments. Phycobilisomes are large light-harvesting protein complexes found in cyanobacteria and red ...Photosynthetic organisms have developed various light-harvesting systems to adapt to their environments. Phycobilisomes are large light-harvesting protein complexes found in cyanobacteria and red algae, although how the energies of the chromophores within these complexes are modulated by their environment is unclear. Here we report the cryo-electron microscopy structure of a 14.7-megadalton phycobilisome with a hemiellipsoidal shape from the red alga Porphyridium purpureum. Within this complex we determine the structures of 706 protein subunits, including 528 phycoerythrin, 72 phycocyanin, 46 allophycocyanin and 60 linker proteins. In addition, 1,598 chromophores are resolved comprising 1,430 phycoerythrobilin, 48 phycourobilin and 120 phycocyanobilin molecules. The markedly improved resolution of our structure compared with that of the phycobilisome of Griffithsia pacifica enabled us to build an accurate atomic model of the P. purpureum phycobilisome system. The model reveals how the linker proteins affect the microenvironment of the chromophores, and suggests that interactions of the aromatic amino acids of the linker proteins with the chromophores may be a key factor in fine-tuning the energy states of the chromophores to ensure the efficient unidirectional transfer of energy. | ||||||||||||
| History |
| ||||||||||||
| Remark 0 | THIS DEPOSITION 7LJ0 PROVIDES A STRUCTURAL MODEL FOR PREVIOUSLY UNMODELED EM MAP DENSITY IN EMD- ...THIS DEPOSITION 7LJ0 PROVIDES A STRUCTURAL MODEL FOR PREVIOUSLY UNMODELED EM MAP DENSITY IN EMD-9976. THIS IS NOT A REPLACEMENT, ONLY AN ADDITION. ORIGINAL AUTHORS OF EMD-9976 ARE SUI, S.F., MA, J.F., YOU, X., SUN, S. |
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7lj0.cif.gz 7lj0.cif.gz | 55.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7lj0.ent.gz pdb7lj0.ent.gz | 34.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7lj0.json.gz 7lj0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lj/7lj0 https://data.pdbj.org/pub/pdb/validation_reports/lj/7lj0 ftp://data.pdbj.org/pub/pdb/validation_reports/lj/7lj0 ftp://data.pdbj.org/pub/pdb/validation_reports/lj/7lj0 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  9976M  7lixC  7liyC  7lizC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 34958.746 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Porphyridium purpureum (eukaryote) / References: UniProt: A0A5J4Z323 Porphyridium purpureum (eukaryote) / References: UniProt: A0A5J4Z323 | ||||
|---|---|---|---|---|---|
| #2: Protein | Mass: 18584.182 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Porphyridium purpureum (eukaryote) / References: UniProt: P11393 Porphyridium purpureum (eukaryote) / References: UniProt: P11393 | ||||
| #3: Chemical | | Has ligand of interest | N | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Phycobilisome / Type: COMPLEX / Entity ID: #1-#2 / Source: NATURAL |
|---|---|
| Source (natural) | Organism:  Porphyridium purpureum (eukaryote) Porphyridium purpureum (eukaryote) |
| Buffer solution | pH: 8 |
| Specimen | Embedding applied: YES / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| EM embedding | Material: ice |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: OTHER FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: OTHER |
| Electron lens | Mode: BRIGHT FIELD |
| Image recording | Electron dose: 48 e/Å2 / Film or detector model: GATAN K2 SUMMIT (4k x 4k) |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||
| CTF correction | Type: PHASE FLIPPING ONLY | ||||||||||||||||||||||||
| 3D reconstruction | Resolution: 2.8 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 191825 / Symmetry type: POINT | ||||||||||||||||||||||||
| Atomic model building | 3D fitting-ID: 1 / Source name: PDB / Type: experimental model
| ||||||||||||||||||||||||
| Refinement | Cross valid method: NONE Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2 | ||||||||||||||||||||||||
| Displacement parameters | Biso mean: 21.75 Å2 | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj



