+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 6z5y | ||||||
|---|---|---|---|---|---|---|---|
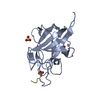
| タイトル | Structure of a novel LPMO from Phytophthora infestans | ||||||
 要素 要素 | Lytic Polysaccharide Monooxygenase | ||||||
 キーワード キーワード | METAL BINDING PROTEIN / Lytic polysaccharide monooxygenase / copper / LPMO | ||||||
| 機能・相同性 | metal ion binding / COPPER (II) ION / TRIETHYLENE GLYCOL / Mucin-like protein 機能・相同性情報 機能・相同性情報 | ||||||
| 生物種 |  Phytophthora infestans (真核生物) Phytophthora infestans (真核生物) | ||||||
| 手法 |  X線回折 / X線回折 /  シンクロトロン / AB INITIO PHASING / 解像度: 1.01 Å シンクロトロン / AB INITIO PHASING / 解像度: 1.01 Å | ||||||
 データ登録者 データ登録者 | Urresti, S. / Davies, G.J. | ||||||
| 資金援助 |  英国, 1件 英国, 1件
| ||||||
 引用 引用 |  ジャーナル: Science / 年: 2021 ジャーナル: Science / 年: 2021タイトル: Secreted pectin monooxygenases drive plant infection by pathogenic oomycetes. 著者: Sabbadin, F. / Urresti, S. / Henrissat, B. / Avrova, A.O. / Welsh, L.R.J. / Lindley, P.J. / Csukai, M. / Squires, J.N. / Walton, P.H. / Davies, G.J. / Bruce, N.C. / Whisson, S.C. / McQueen-Mason, S.J. | ||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  6z5y.cif.gz 6z5y.cif.gz | 167.2 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb6z5y.ent.gz pdb6z5y.ent.gz | 131.8 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  6z5y.json.gz 6z5y.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| 文書・要旨 |  6z5y_validation.pdf.gz 6z5y_validation.pdf.gz | 264 KB | 表示 |  wwPDB検証レポート wwPDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  6z5y_full_validation.pdf.gz 6z5y_full_validation.pdf.gz | 267.6 KB | 表示 | |
| XML形式データ |  6z5y_validation.xml.gz 6z5y_validation.xml.gz | 9.4 KB | 表示 | |
| CIF形式データ |  6z5y_validation.cif.gz 6z5y_validation.cif.gz | 15.7 KB | 表示 | |
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/z5/6z5y https://data.pdbj.org/pub/pdb/validation_reports/z5/6z5y ftp://data.pdbj.org/pub/pdb/validation_reports/z5/6z5y ftp://data.pdbj.org/pub/pdb/validation_reports/z5/6z5y | HTTPS FTP |
-関連構造データ
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 単位格子 |
|
- 要素
要素
| #1: タンパク質 | 分子量: 20016.531 Da / 分子数: 2 / 由来タイプ: 組換発現 / 詳細: C-terminus Strep tag sequence: WSHPQFEK 由来: (組換発現)  Phytophthora infestans (strain T30-4) (真核生物) Phytophthora infestans (strain T30-4) (真核生物)遺伝子: PITG_04949 / 発現宿主:  #2: 化合物 | #3: 化合物 | ChemComp-PGE / | #4: 水 | ChemComp-HOH / | 研究の焦点であるリガンドがあるか | Y | Has protein modification | Y | |
|---|
-実験情報
-実験
| 実験 | 手法:  X線回折 / 使用した結晶の数: 1 X線回折 / 使用した結晶の数: 1 |
|---|
- 試料調製
試料調製
| 結晶 | マシュー密度: 2.13 Å3/Da / 溶媒含有率: 42.18 % / 解説: Hollow rods |
|---|---|
| 結晶化 | 温度: 293 K / 手法: 蒸気拡散法, シッティングドロップ法 / pH: 7 詳細: 20mM Tris pH 7.0, 22% Polyethylene Glycol (PEG) 3350 |
-データ収集
| 回折 | 平均測定温度: 100 K / Serial crystal experiment: N | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 放射光源 | 由来:  シンクロトロン / サイト: シンクロトロン / サイト:  Diamond Diamond  / ビームライン: I04 / 波長: 0.916 Å / ビームライン: I04 / 波長: 0.916 Å | ||||||||||||||||||||||||
| 検出器 | タイプ: DECTRIS PILATUS 6M-F / 検出器: PIXEL / 日付: 2018年12月1日 | ||||||||||||||||||||||||
| 放射 | プロトコル: SINGLE WAVELENGTH / 単色(M)・ラウエ(L): M / 散乱光タイプ: x-ray | ||||||||||||||||||||||||
| 放射波長 | 波長: 0.916 Å / 相対比: 1 | ||||||||||||||||||||||||
| 反射 | 解像度: 1.01→28.79 Å / Num. obs: 156702 / % possible obs: 92.4 % / 冗長度: 3.3 % / CC1/2: 0.998 / Rmerge(I) obs: 0.048 / Rpim(I) all: 0.03 / Rrim(I) all: 0.057 / Net I/σ(I): 8.9 | ||||||||||||||||||||||||
| 反射 シェル | Diffraction-ID: 1
|
- 解析
解析
| ソフトウェア |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 構造決定の手法: AB INITIO PHASING / 解像度: 1.01→28.79 Å / Cor.coef. Fo:Fc: 0.978 / Cor.coef. Fo:Fc free: 0.973 / SU B: 0.791 / SU ML: 0.018 / 交差検証法: THROUGHOUT / σ(F): 0 / ESU R: 0.025 / ESU R Free: 0.025 / 立体化学のターゲット値: MAXIMUM LIKELIHOOD 詳細: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY Residues 139-142 and 172-177 (chains A and B) show poor density and therefore low occupancy. C-terminal ...詳細: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY Residues 139-142 and 172-177 (chains A and B) show poor density and therefore low occupancy. C-terminal residues (172-177) correspond to a non cleaved Strep tag, and some have been built with no side chains.
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 溶媒の処理 | イオンプローブ半径: 0.8 Å / 減衰半径: 0.8 Å / VDWプローブ半径: 1.2 Å / 溶媒モデル: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 原子変位パラメータ | Biso max: 36.26 Å2 / Biso mean: 11.889 Å2 / Biso min: 5.72 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 精密化ステップ | サイクル: final / 解像度: 1.01→28.79 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 拘束条件 |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS精密化 シェル | 解像度: 1.01→1.036 Å / Rfactor Rfree error: 0 / Total num. of bins used: 20
|
 ムービー
ムービー コントローラー
コントローラー











 PDBj
PDBj




