+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6s2i | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
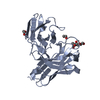
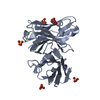
| Title | Anti-tumor antibody 14F7-derived scFv in complex with NeuGc Gm3 | |||||||||
 Components Components | scFv C1 | |||||||||
 Keywords Keywords | ANTITUMOR PROTEIN / ganglioside / complex / scfv / antibody fragment / single chain fragment variable / immuno therapy / 14F7 / gm3 | |||||||||
| Biological species |  | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.285 Å MOLECULAR REPLACEMENT / Resolution: 2.285 Å | |||||||||
 Authors Authors | Bjerregaard-Andersen, K. / Heggelund, J.E. / Krengel, U. | |||||||||
 Citation Citation |  Journal: Glycobiology / Year: 2021 Journal: Glycobiology / Year: 2021Title: Key role of a structural water molecule for the specificity of 14F7-An antitumor antibody targeting the NeuGc GM3 ganglioside. Authors: Bjerregaard-Andersen, K. / Abraha, F. / Johannesen, H. / Oscarson, S. / Moreno, E. / Krengel, U. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6s2i.cif.gz 6s2i.cif.gz | 208.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6s2i.ent.gz pdb6s2i.ent.gz | 157.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6s2i.json.gz 6s2i.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/s2/6s2i https://data.pdbj.org/pub/pdb/validation_reports/s2/6s2i ftp://data.pdbj.org/pub/pdb/validation_reports/s2/6s2i ftp://data.pdbj.org/pub/pdb/validation_reports/s2/6s2i | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6ffjS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Antibody | Mass: 28116.238 Da / Num. of mol.: 8 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Polysaccharide | N-glycolyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | Type: oligosaccharide / Mass: 649.551 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source #3: Water | ChemComp-HOH / | Has ligand of interest | N | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.33 Å3/Da / Density % sol: 47.11 % / Description: Plates |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 8.5 Details: 12.5 % w/v PEG 1000, 12.5 % w/v PEG 3350, 12.5 % v/v MPD, 0.02 M 1,6-hexandiol, 0.02 M 1-butanol, 0.02 M (RS)-1,2-propanediol, 0.02 M 2-propanol, 0.02 M 1,4-butanediol, 0.02 M 1,3- ...Details: 12.5 % w/v PEG 1000, 12.5 % w/v PEG 3350, 12.5 % v/v MPD, 0.02 M 1,6-hexandiol, 0.02 M 1-butanol, 0.02 M (RS)-1,2-propanediol, 0.02 M 2-propanol, 0.02 M 1,4-butanediol, 0.02 M 1,3-propanediol, 0.1 M bicine/tris base pH 8.5 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: MASSIF-3 / Wavelength: 0.9677 Å / Beamline: MASSIF-3 / Wavelength: 0.9677 Å |
| Detector | Type: DECTRIS EIGER X 4M / Detector: PIXEL / Date: May 15, 2017 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9677 Å / Relative weight: 1 |
| Reflection | Resolution: 2.285→62.9 Å / Num. obs: 42116 / % possible obs: 98.8 % / Redundancy: 4.2 % / Biso Wilson estimate: 44.029 Å2 / CC1/2: 0.98 / Rmerge(I) obs: 0.097 / Rpim(I) all: 0.053 / Rrim(I) all: 0.111 / Net I/σ(I): 9.03 |
| Reflection shell | Resolution: 2.285→2.325 Å / Redundancy: 4.4 % / Rmerge(I) obs: 0.697 / Mean I/σ(I) obs: 2.1 / Num. unique obs: 2141 / CC1/2: 0.813 / Rpim(I) all: 0.379 / Rrim(I) all: 0.795 / % possible all: 99.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 6FFJ Resolution: 2.285→62.865 Å / SU ML: 0.35 / Cross valid method: FREE R-VALUE / σ(F): 1.35 / Phase error: 36.43
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 51.46 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.285→62.865 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj


