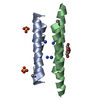
Entry Database : PDB / ID : 6q5qTitle Crystal structure of a CC-Hex mutant that forms an antiparallel four-helix coiled coil CC-Hex-KgEb CC-Hex-KgEb Keywords / / / / Function / homology Biological species synthetic construct (others) Method / / / Resolution : 1.08 Å Authors Zaccai, N.R. / Rhys, G.G. / Wood, C.W. / Beesley, J.L. / Brady, R.L. / Woolfson, D.N. Funding support Organization Grant number Country Biotechnology and Biological Sciences Research Council BB/J014400/1 Engineering and Physical Sciences Research Council EP/G036764/1 European Research Council 340764 Biotechnology and Biological Sciences Research Council BB/R00661X/1 Engineering and Physical Sciences Research Council EP/K03927X/1
Journal : J.Am.Chem.Soc. / Year : 2019Title : Navigating the Structural Landscape of De Novo alpha-Helical Bundles.Authors : Rhys, G.G. / Wood, C.W. / Beesley, J.L. / Zaccai, N.R. / Burton, A.J. / Brady, R.L. / Thomson, A.R. / Woolfson, D.N. History Deposition Dec 9, 2018 Deposition site / Processing site Revision 1.0 May 22, 2019 Provider / Type Revision 1.1 Jun 19, 2019 Group / Database references / Category / citation_author / pdbx_database_procItem _citation.journal_volume / _citation.page_first ... _citation.journal_volume / _citation.page_first / _citation.page_last / _citation.title / _citation_author.identifier_ORCID / _citation_author.name Revision 1.2 Nov 6, 2024 Group / Database references / Structure summaryCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_entry_details / pdbx_modification_feature Item / _database_2.pdbx_database_accession
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON / AB INITIO PHASING / Resolution: 1.08 Å
SYNCHROTRON / AB INITIO PHASING / Resolution: 1.08 Å  Authors
Authors United Kingdom,
United Kingdom,  Belgium, 5items
Belgium, 5items  Citation
Citation Journal: J.Am.Chem.Soc. / Year: 2019
Journal: J.Am.Chem.Soc. / Year: 2019 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6q5q.cif.gz
6q5q.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6q5q.ent.gz
pdb6q5q.ent.gz PDB format
PDB format 6q5q.json.gz
6q5q.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/q5/6q5q
https://data.pdbj.org/pub/pdb/validation_reports/q5/6q5q ftp://data.pdbj.org/pub/pdb/validation_reports/q5/6q5q
ftp://data.pdbj.org/pub/pdb/validation_reports/q5/6q5q









 Links
Links Assembly
Assembly

 Components
Components X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  Diamond
Diamond  / Beamline: I02 / Wavelength: 0.9795 Å
/ Beamline: I02 / Wavelength: 0.9795 Å Processing
Processing Movie
Movie Controller
Controller










 PDBj
PDBj



