[English] 日本語
 Yorodumi
Yorodumi- PDB-6g9l: Structure of homomeric mLRRC8A volume-regulated anion channel at ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6g9l | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of homomeric mLRRC8A volume-regulated anion channel at 5.01 A resolution | ||||||
 Components Components | Volume-regulated anion channel subunit LRRC8A | ||||||
 Keywords Keywords | MEMBRANE PROTEIN / Chloride channel / Swelling-activated / VSOAC / Leucine-rich repeat | ||||||
| Function / homology |  Function and homology information Function and homology informationMiscellaneous transport and binding events / pre-B cell differentiation / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / monoatomic anion transport / cell volume homeostasis ...Miscellaneous transport and binding events / pre-B cell differentiation / volume-sensitive anion channel activity / aspartate transmembrane transport / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / monoatomic anion transport / cell volume homeostasis / protein hexamerization / response to osmotic stress / monoatomic ion channel complex / positive regulation of myoblast differentiation / intracellular glucose homeostasis / chloride transmembrane transport / positive regulation of insulin secretion / spermatogenesis / lysosomal membrane / cell surface / membrane / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 5.01 Å | ||||||
 Authors Authors | Sawicka, M. / Deneka, D. / Lam, A.K.M. / Paulino, C. / Dutzler, R. | ||||||
| Funding support |  Switzerland, 1items Switzerland, 1items
| ||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Structure of a volume-regulated anion channel of the LRRC8 family. Authors: Dawid Deneka / Marta Sawicka / Andy K M Lam / Cristina Paulino / Raimund Dutzler /   Abstract: Volume-regulated anion channels are activated in response to hypotonic stress. These channels are composed of closely related paralogues of the leucine-rich repeat-containing protein 8 (LRRC8) family ...Volume-regulated anion channels are activated in response to hypotonic stress. These channels are composed of closely related paralogues of the leucine-rich repeat-containing protein 8 (LRRC8) family that co-assemble to form hexameric complexes. Here, using cryo-electron microscopy and X-ray crystallography, we determine the structure of a homomeric channel of the obligatory subunit LRRC8A. This protein conducts ions and has properties in common with endogenous heteromeric channels. Its modular structure consists of a transmembrane pore domain followed by a cytoplasmic leucine-rich repeat domain. The transmembrane domain, which is structurally related to connexin proteins, is wide towards the cytoplasm but constricted on the outside by a structural unit that acts as a selectivity filter. An excess of basic residues in the filter and throughout the pore attracts anions by electrostatic interaction. Our work reveals the previously unknown architecture of volume-regulated anion channels and their mechanism of selective anion conduction. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6g9l.cif.gz 6g9l.cif.gz | 754.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6g9l.ent.gz pdb6g9l.ent.gz | 634.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6g9l.json.gz 6g9l.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/g9/6g9l https://data.pdbj.org/pub/pdb/validation_reports/g9/6g9l ftp://data.pdbj.org/pub/pdb/validation_reports/g9/6g9l ftp://data.pdbj.org/pub/pdb/validation_reports/g9/6g9l | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4366MC  4361C  4362C  4367C 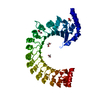 6fnwC  6g8zC  6g9oC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 94239.383 Da / Num. of mol.: 6 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / References: UniProt: Q80WG5 Homo sapiens (human) / References: UniProt: Q80WG5Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Homohexameric mLRRC8A / Type: COMPLEX / Entity ID: all / Source: RECOMBINANT | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Source (natural) | Organism:  | ||||||||||||||||||||
| Source (recombinant) | Organism:  Homo sapiens (human) / Cell: HEK293S-GnTI minus Homo sapiens (human) / Cell: HEK293S-GnTI minus | ||||||||||||||||||||
| Buffer solution | pH: 8.5 | ||||||||||||||||||||
| Buffer component |
| ||||||||||||||||||||
| Specimen | Conc.: 3.9 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES | ||||||||||||||||||||
| Specimen support | Grid material: GOLD / Grid mesh size: 200 divisions/in. / Grid type: Quantifoil R1.2/1.3 | ||||||||||||||||||||
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI POLARA 300 |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 37313 X / Calibrated magnification: 37313 X / Nominal defocus max: 3200 nm / Nominal defocus min: 500 nm / Cs: 2 mm |
| Specimen holder | Cryogen: NITROGEN |
| Image recording | Average exposure time: 12.5 sec. / Electron dose: 1.2 e/Å2 / Detector mode: SUPER-RESOLUTION / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Num. of real images: 9209 |
| Image scans | Movie frames/image: 50 / Used frames/image: 1-50 |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.13_2998: / Classification: refinement | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||||||||
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 329624 | ||||||||||||||||||||||||||||||
| Symmetry | Point symmetry: C3 (3 fold cyclic) | ||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 5.01 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 54861 / Symmetry type: POINT | ||||||||||||||||||||||||||||||
| Atomic model building | Protocol: RIGID BODY FIT | ||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller









 PDBj
PDBj




