[English] 日本語
 Yorodumi
Yorodumi- PDB-6fnw: Structure of a volume-regulated anion channel of the LRRC8 family -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6fnw | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of a volume-regulated anion channel of the LRRC8 family | ||||||
 Components Components | Volume-regulated anion channel subunit LRRC8A | ||||||
 Keywords Keywords | MEMBRANE PROTEIN / ion channel / leucine-rich repeat domain | ||||||
| Function / homology |  Function and homology information Function and homology informationMiscellaneous transport and binding events / pre-B cell differentiation / aspartate transmembrane transport / volume-sensitive anion channel activity / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / protein hexamerization / monoatomic anion transport ...Miscellaneous transport and binding events / pre-B cell differentiation / aspartate transmembrane transport / volume-sensitive anion channel activity / cyclic-GMP-AMP transmembrane transporter activity / cyclic-GMP-AMP transmembrane import across plasma membrane / taurine transmembrane transport / monoatomic anion transmembrane transport / protein hexamerization / monoatomic anion transport / cell volume homeostasis / response to osmotic stress / intracellular glucose homeostasis / monoatomic ion channel complex / positive regulation of myoblast differentiation / chloride transmembrane transport / positive regulation of insulin secretion / spermatogenesis / lysosomal membrane / cell surface / identical protein binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.8 Å SAD / Resolution: 1.8 Å | ||||||
 Authors Authors | Deneka, D. / Sawicka, M. / Lam, A.K.M. / Paulino, C. / Dutzler, R. | ||||||
| Funding support |  Switzerland, 1items Switzerland, 1items
| ||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Structure of a volume-regulated anion channel of the LRRC8 family. Authors: Dawid Deneka / Marta Sawicka / Andy K M Lam / Cristina Paulino / Raimund Dutzler /   Abstract: Volume-regulated anion channels are activated in response to hypotonic stress. These channels are composed of closely related paralogues of the leucine-rich repeat-containing protein 8 (LRRC8) family ...Volume-regulated anion channels are activated in response to hypotonic stress. These channels are composed of closely related paralogues of the leucine-rich repeat-containing protein 8 (LRRC8) family that co-assemble to form hexameric complexes. Here, using cryo-electron microscopy and X-ray crystallography, we determine the structure of a homomeric channel of the obligatory subunit LRRC8A. This protein conducts ions and has properties in common with endogenous heteromeric channels. Its modular structure consists of a transmembrane pore domain followed by a cytoplasmic leucine-rich repeat domain. The transmembrane domain, which is structurally related to connexin proteins, is wide towards the cytoplasm but constricted on the outside by a structural unit that acts as a selectivity filter. An excess of basic residues in the filter and throughout the pore attracts anions by electrostatic interaction. Our work reveals the previously unknown architecture of volume-regulated anion channels and their mechanism of selective anion conduction. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6fnw.cif.gz 6fnw.cif.gz | 278.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6fnw.ent.gz pdb6fnw.ent.gz | 230.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6fnw.json.gz 6fnw.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/fn/6fnw https://data.pdbj.org/pub/pdb/validation_reports/fn/6fnw ftp://data.pdbj.org/pub/pdb/validation_reports/fn/6fnw ftp://data.pdbj.org/pub/pdb/validation_reports/fn/6fnw | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4361C  4362C  4366C  4367C  6g8zC  6g9lC  6g9oC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 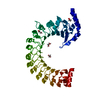
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 48155.094 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Details: C-terminal domain of mouse LRRC8A / Source: (gene. exp.)   Homo sapiens (human) / References: UniProt: Q80WG5 Homo sapiens (human) / References: UniProt: Q80WG5 | ||
|---|---|---|---|
| #2: Chemical | ChemComp-EDO / #3: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.49 Å3/Da / Density % sol: 50.56 % |
|---|---|
| Crystal grow | Temperature: 277.15 K / Method: vapor diffusion, sitting drop / pH: 8.5 Details: 22% PEG 3350, 0.2M Na/K tartrate, 0.1M Bis-Tris propane pH 8.5 |
-Data collection
| Diffraction | Mean temperature: 80 K | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SLS SLS  / Beamline: X06DA / Wavelength: 1, 2.05 / Beamline: X06DA / Wavelength: 1, 2.05 | |||||||||
| Detector | Type: DECTRIS PILATUS 2M-F / Detector: PIXEL / Date: Nov 18, 2016 | |||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | |||||||||
| Radiation wavelength |
| |||||||||
| Reflection | Resolution: 1.8→40.452 Å / Num. obs: 44375 / % possible obs: 100 % / Redundancy: 20.2 % / Net I/σ(I): 27.05 | |||||||||
| Reflection shell | Resolution: 1.8→1.9 Å / Redundancy: 19.8 % / Mean I/σ(I) obs: 2.27 / Num. unique obs: 6557 / CC1/2: 0.9 / Rrim(I) all: 1.58 / % possible all: 99.9 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD / Resolution: 1.8→40.452 Å / SU ML: 0.22 / Cross valid method: FREE R-VALUE / σ(F): 1.35 / Phase error: 26.35 SAD / Resolution: 1.8→40.452 Å / SU ML: 0.22 / Cross valid method: FREE R-VALUE / σ(F): 1.35 / Phase error: 26.35
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→40.452 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 16.738 Å / Origin y: -45.7676 Å / Origin z: -1.9617 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: all |
 Movie
Movie Controller
Controller










 PDBj
PDBj







