[English] 日本語
 Yorodumi
Yorodumi- PDB-6dn3: CRYSTAL STRUCTURE OF THE FMN RIBOSWITCH BOUND TO BRX1555 SPLIT RNA -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6dn3 | ||||||
|---|---|---|---|---|---|---|---|
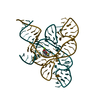
| Title | CRYSTAL STRUCTURE OF THE FMN RIBOSWITCH BOUND TO BRX1555 SPLIT RNA | ||||||
 Components Components |
| ||||||
 Keywords Keywords | RNA / FMN / RIBOSWITCH / TRANSCRIPTION | ||||||
| Function / homology | Chem-GZ4 / : / RNA / RNA (> 10) Function and homology information Function and homology information | ||||||
| Biological species |  Fusobacterium nucleatum (bacteria) Fusobacterium nucleatum (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.8 Å MOLECULAR REPLACEMENT / Resolution: 2.8 Å | ||||||
 Authors Authors | Vicens, Q. / Mondragon, E. / Reyes, F.E. / Berman, J. / Kaur, H. / Kells, K. / Wickens, P. / Wilson, J. / Gadwood, R. / Schostarez, H. ...Vicens, Q. / Mondragon, E. / Reyes, F.E. / Berman, J. / Kaur, H. / Kells, K. / Wickens, P. / Wilson, J. / Gadwood, R. / Schostarez, H. / Suto, R.K. / Coish, P. / Blount, K.F. / Batey, R.T. | ||||||
 Citation Citation |  Journal: ACS Chem. Biol. / Year: 2018 Journal: ACS Chem. Biol. / Year: 2018Title: Structure-Activity Relationship of Flavin Analogues That Target the Flavin Mononucleotide Riboswitch. Authors: Vicens, Q. / Mondragon, E. / Reyes, F.E. / Coish, P. / Aristoff, P. / Berman, J. / Kaur, H. / Kells, K.W. / Wickens, P. / Wilson, J. / Gadwood, R.C. / Schostarez, H.J. / Suto, R.K. / Blount, K.F. / Batey, R.T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6dn3.cif.gz 6dn3.cif.gz | 73.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6dn3.ent.gz pdb6dn3.ent.gz | 52.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6dn3.json.gz 6dn3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/dn/6dn3 https://data.pdbj.org/pub/pdb/validation_reports/dn/6dn3 ftp://data.pdbj.org/pub/pdb/validation_reports/dn/6dn3 ftp://data.pdbj.org/pub/pdb/validation_reports/dn/6dn3 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6dn1C  6dn2C  3f4eS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-RNA chain , 2 types, 2 molecules XY
| #1: RNA chain | Mass: 17455.389 Da / Num. of mol.: 1 / Source method: obtained synthetically Details: IN VITRO TRANSCRIBED RNA FROM FUSOBACTERIUM NUCLEATUM IMPX GENE Source: (synth.)  Fusobacterium nucleatum (bacteria) Fusobacterium nucleatum (bacteria) |
|---|---|
| #2: RNA chain | Mass: 18092.771 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  Fusobacterium nucleatum (bacteria) Fusobacterium nucleatum (bacteria) |
-Non-polymers , 5 types, 14 molecules 








| #3: Chemical | ChemComp-CL / | ||||||
|---|---|---|---|---|---|---|---|
| #4: Chemical | ChemComp-MG / #5: Chemical | ChemComp-GZ4 / | #6: Chemical | ChemComp-K / | #7: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.92 Å3/Da / Density % sol: 57.81 % |
|---|---|
| Crystal grow | Temperature: 293.15 K / Method: evaporation / pH: 6.5 / Details: PEG 4000 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X29A / Wavelength: 1.075 Å / Beamline: X29A / Wavelength: 1.075 Å |
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Sep 21, 2009 / Details: MIRRORS |
| Radiation | Monochromator: SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.075 Å / Relative weight: 1 |
| Reflection | Resolution: 2.8→25 Å / Num. obs: 10406 / % possible obs: 96.8 % / Observed criterion σ(I): 0 / Redundancy: 11.2 % / Biso Wilson estimate: 61.65 Å2 / Rmerge(I) obs: 0.074 / Net I/σ(I): 24.6 |
| Reflection shell | Resolution: 2.8→2.9 Å / Redundancy: 2.7 % / Rmerge(I) obs: 0.378 / Mean I/σ(I) obs: 2.3 / % possible all: 87 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3F4E Resolution: 2.8→23.1 Å / Cor.coef. Fo:Fc: 0.94 / Cor.coef. Fo:Fc free: 0.91 / Rfactor Rfree error: 0 / SU R Cruickshank DPI: 0.92 / Cross valid method: THROUGHOUT / σ(F): 0 / SU Rfree Blow DPI: 0.32 / SU Rfree Cruickshank DPI: 0.302
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 147.87 Å2 / Biso mean: 81.86 Å2 / Biso min: 45.24 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.37 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.8→23.1 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.8→3.13 Å / Total num. of bins used: 5
|
 Movie
Movie Controller
Controller











 PDBj
PDBj

































