[English] 日本語
 Yorodumi
Yorodumi- PDB-6bx8: Human Mesotrypsin (PRSS3) Complexed with Tissue Factor Pathway In... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6bx8 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
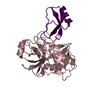
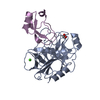
| Title | Human Mesotrypsin (PRSS3) Complexed with Tissue Factor Pathway Inhibitor Variant (TFPI1-KD1-K15R-I17C-I34C) | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | HYDROLASE / Hydrolase Inhibitor | |||||||||
| Function / homology |  Function and homology information Function and homology informationUptake of dietary cobalamins into enterocytes / antimicrobial humoral response / Alpha-defensins / Differentiation of Keratinocytes in Interfollicular Epidermis in Mammalian Skin / : / Antimicrobial peptides / zymogen activation / cellular response to steroid hormone stimulus / endopeptidase inhibitor activity / trypsin ...Uptake of dietary cobalamins into enterocytes / antimicrobial humoral response / Alpha-defensins / Differentiation of Keratinocytes in Interfollicular Epidermis in Mammalian Skin / : / Antimicrobial peptides / zymogen activation / cellular response to steroid hormone stimulus / endopeptidase inhibitor activity / trypsin / negative regulation of blood coagulation / endothelial cell migration / digestion / side of membrane / serine-type peptidase activity / serine-type endopeptidase inhibitor activity / caveola / blood coagulation / tertiary granule lumen / serine-type endopeptidase activity / calcium ion binding / Neutrophil degranulation / cell surface / proteolysis / : / extracellular region / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.98 Å molecular replacement / Resolution: 1.98 Å | |||||||||
 Authors Authors | Coban, M. / Sankaran, B. / Cohen, I. / Hockla, A. / Papo, N. / Radisky, E.S. | |||||||||
| Funding support |  United States, United States,  Israel, 2items Israel, 2items
| |||||||||
 Citation Citation |  Journal: J. Biol. Chem. / Year: 2019 Journal: J. Biol. Chem. / Year: 2019Title: Disulfide engineering of human Kunitz-type serine protease inhibitors enhances proteolytic stability and target affinity toward mesotrypsin. Authors: Cohen, I. / Coban, M. / Shahar, A. / Sankaran, B. / Hockla, A. / Lacham, S. / Caulfield, T.R. / Radisky, E.S. / Papo, N. #1: Journal: Acta Crystallogr. D Biol. Crystallogr. / Year: 2008 Title: Model preparation in MOLREP and examples of model improvement using X-ray data. Authors: Lebedev, A.A. / Vagin, A.A. / Murshudov, G.N. #2: Journal: Acta Crystallogr. D Biol. Crystallogr. / Year: 1997 Title: Refinement of macromolecular structures by the maximum-likelihood method. Authors: Murshudov, G.N. / Vagin, A.A. / Dodson, E.J. #3: Journal: Acta Crystallogr D Biol Crystallogr / Year: 2004 Title: Coot: model-building tools for molecular graphics. Authors: Paul Emsley / Kevin Cowtan /  Abstract: CCP4mg is a project that aims to provide a general-purpose tool for structural biologists, providing tools for X-ray structure solution, structure comparison and analysis, and publication-quality ...CCP4mg is a project that aims to provide a general-purpose tool for structural biologists, providing tools for X-ray structure solution, structure comparison and analysis, and publication-quality graphics. The map-fitting tools are available as a stand-alone package, distributed as 'Coot'. #4: Journal: Acta Crystallogr D Biol Crystallogr / Year: 2010 Title: PHENIX: a comprehensive Python-based system for macromolecular structure solution. Authors: Paul D Adams / Pavel V Afonine / Gábor Bunkóczi / Vincent B Chen / Ian W Davis / Nathaniel Echols / Jeffrey J Headd / Li-Wei Hung / Gary J Kapral / Ralf W Grosse-Kunstleve / Airlie J McCoy ...Authors: Paul D Adams / Pavel V Afonine / Gábor Bunkóczi / Vincent B Chen / Ian W Davis / Nathaniel Echols / Jeffrey J Headd / Li-Wei Hung / Gary J Kapral / Ralf W Grosse-Kunstleve / Airlie J McCoy / Nigel W Moriarty / Robert Oeffner / Randy J Read / David C Richardson / Jane S Richardson / Thomas C Terwilliger / Peter H Zwart /  Abstract: Macromolecular X-ray crystallography is routinely applied to understand biological processes at a molecular level. However, significant time and effort are still required to solve and complete many ...Macromolecular X-ray crystallography is routinely applied to understand biological processes at a molecular level. However, significant time and effort are still required to solve and complete many of these structures because of the need for manual interpretation of complex numerical data using many software packages and the repeated use of interactive three-dimensional graphics. PHENIX has been developed to provide a comprehensive system for macromolecular crystallographic structure solution with an emphasis on the automation of all procedures. This has relied on the development of algorithms that minimize or eliminate subjective input, the development of algorithms that automate procedures that are traditionally performed by hand and, finally, the development of a framework that allows a tight integration between the algorithms. #5: Journal: Acta Crystallogr. D Biol. Crystallogr. / Year: 2010 Title: MolProbity: all-atom structure validation for macromolecular crystallography. Authors: Chen, V.B. / Arendall, W.B. / Headd, J.J. / Keedy, D.A. / Immormino, R.M. / Kapral, G.J. / Murray, L.W. / Richardson, J.S. / Richardson, D.C. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6bx8.cif.gz 6bx8.cif.gz | 610.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6bx8.ent.gz pdb6bx8.ent.gz | 513.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6bx8.json.gz 6bx8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bx/6bx8 https://data.pdbj.org/pub/pdb/validation_reports/bx/6bx8 ftp://data.pdbj.org/pub/pdb/validation_reports/bx/6bx8 ftp://data.pdbj.org/pub/pdb/validation_reports/bx/6bx8 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 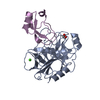 6harC  3p95S  4bqdS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 24257.457 Da / Num. of mol.: 4 / Mutation: S257A Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PRSS3, PRSS4, TRY3, TRY4 / Production host: Homo sapiens (human) / Gene: PRSS3, PRSS4, TRY3, TRY4 / Production host:  #2: Protein | Mass: 9681.825 Da / Num. of mol.: 4 / Mutation: K64R, M66C, I83C Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: TFPI, LACI, TFPI1 / Production host: Homo sapiens (human) / Gene: TFPI, LACI, TFPI1 / Production host:  Komagataella pastoris (fungus) / References: UniProt: P10646 Komagataella pastoris (fungus) / References: UniProt: P10646#3: Chemical | ChemComp-SO4 / #4: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.04 Å3/Da / Density % sol: 39.64 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 4.6 Details: 0.2M Lithium Sulfate, 0.1M BIS-Tris-HCl pH 4.6, 25% v/v PEG 3350 (Anatrace Top96-27) |
-Data collection
| Diffraction | Mean temperature: 80 K / Ambient temp details: LN2 |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ALS ALS  / Beamline: 5.0.1 / Wavelength: 0.97741 Å / Beamline: 5.0.1 / Wavelength: 0.97741 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Aug 2, 2017 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97741 Å / Relative weight: 1 |
| Reflection | Resolution: 1.98→60.0901 Å / Num. obs: 70450 / % possible obs: 97.4 % / Redundancy: 1.7 % / Biso Wilson estimate: 19.36 Å2 / CC1/2: 0.97 / Rmerge(I) obs: 0.067 / Rpim(I) all: 0.067 / Rrim(I) all: 0.095 / Net I/σ(I): 6.5 |
| Reflection shell | Resolution: 1.98→2.05 Å / Redundancy: 1.7 % / Rmerge(I) obs: 0.249 / Mean I/σ(I) obs: 2.5 / Num. unique obs: 7014 / CC1/2: 0.887 / Rpim(I) all: 0.249 / Rrim(I) all: 0.352 / % possible all: 93.7 |
-Phasing
| Phasing | Method:  molecular replacement molecular replacement | ||||||
|---|---|---|---|---|---|---|---|
| Phasing MR | R rigid body: 0.58
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3P95, 4BQD Resolution: 1.98→60.062 Å / SU ML: 0.21 / Cross valid method: THROUGHOUT / σ(F): 1.96 / Phase error: 22.69 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 92.38 Å2 / Biso mean: 30.0686 Å2 / Biso min: 9.88 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.98→60.062 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Rfactor Rfree error: 0 / Total num. of bins used: 24
|
 Movie
Movie Controller
Controller










 PDBj
PDBj












