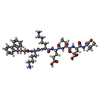
| Entry | Database: PDB / ID: 5d13
|
|---|
| Title | Third PDZ domain (PDZ3) of PSD-95 complexed with CFMOC-KKETEV peptide |
|---|
 Components Components | - CFMOC-KKETEV peptide
- Disks large homolog 4
|
|---|
 Keywords Keywords | signaling protein/inhibitor / protein-peptide complex / signaling protein-inhibitor complex |
|---|
| Function / homology |  Function and homology information Function and homology information
RHO GTPases activate CIT / neuronal ion channel clustering / positive regulation of AMPA glutamate receptor clustering / P2Y1 nucleotide receptor binding / beta-1 adrenergic receptor binding / Neurexins and neuroligins / neuroligin family protein binding / regulation of grooming behavior / structural constituent of postsynaptic density / synaptic vesicle maturation ...RHO GTPases activate CIT / neuronal ion channel clustering / positive regulation of AMPA glutamate receptor clustering / P2Y1 nucleotide receptor binding / beta-1 adrenergic receptor binding / Neurexins and neuroligins / neuroligin family protein binding / regulation of grooming behavior / structural constituent of postsynaptic density / synaptic vesicle maturation / positive regulation of neuron projection arborization / receptor localization to synapse / vocalization behavior / neuron spine / cerebellar mossy fiber / proximal dendrite / protein localization to synapse / LGI-ADAM interactions / Trafficking of AMPA receptors / dendritic spine morphogenesis / negative regulation of receptor internalization / dendritic branch / Activation of Ca-permeable Kainate Receptor / juxtaparanode region of axon / frizzled binding / acetylcholine receptor binding / positive regulation of dendrite morphogenesis / regulation of NMDA receptor activity / cellular response to potassium ion / dendritic spine organization / positive regulation of synapse assembly / RAF/MAP kinase cascade / beta-2 adrenergic receptor binding / NMDA selective glutamate receptor signaling pathway / Synaptic adhesion-like molecules / neuromuscular process controlling balance / neurotransmitter receptor localization to postsynaptic specialization membrane / cortical cytoskeleton / neuron projection terminus / AMPA glutamate receptor clustering / locomotory exploration behavior / AMPA glutamate receptor complex / regulation of neuronal synaptic plasticity / kinesin binding / social behavior / Unblocking of NMDA receptors, glutamate binding and activation / glutamate receptor binding / D1 dopamine receptor binding / positive regulation of synaptic transmission / regulation of postsynaptic membrane neurotransmitter receptor levels / excitatory synapse / ionotropic glutamate receptor binding / dendrite cytoplasm / synaptic membrane / positive regulation of excitatory postsynaptic potential / cell periphery / PDZ domain binding / neuromuscular junction / establishment of protein localization / cell-cell adhesion / regulation of long-term neuronal synaptic plasticity / cerebral cortex development / postsynaptic density membrane / kinase binding / synaptic vesicle / cell-cell junction / cell junction / nervous system development / positive regulation of cytosolic calcium ion concentration / protein-containing complex assembly / scaffold protein binding / protein phosphatase binding / dendritic spine / chemical synaptic transmission / postsynaptic membrane / neuron projection / postsynapse / postsynaptic density / signaling receptor binding / synapse / dendrite / protein kinase binding / protein-containing complex binding / glutamatergic synapse / endoplasmic reticulum / membrane / plasma membrane / cytoplasm / cytosolSimilarity search - Function Polyubiquitination (PEST) N-terminal domain of MAGUK / Disks large homologue 1, N-terminal PEST domain / Polyubiquitination (PEST) N-terminal domain of MAGUK / PDZ-associated domain of NMDA receptors / PDZ-associated domain of NMDA receptors / Disks large 1-like / : / Guanylate kinase, conserved site / Guanylate kinase-like signature. / Guanylate kinase-like domain profile. ...Polyubiquitination (PEST) N-terminal domain of MAGUK / Disks large homologue 1, N-terminal PEST domain / Polyubiquitination (PEST) N-terminal domain of MAGUK / PDZ-associated domain of NMDA receptors / PDZ-associated domain of NMDA receptors / Disks large 1-like / : / Guanylate kinase, conserved site / Guanylate kinase-like signature. / Guanylate kinase-like domain profile. / Guanylate kinase-like domain / Guanylate kinase/L-type calcium channel beta subunit / Guanylate kinase / Guanylate kinase homologues. / PDZ domain / Pdz3 Domain / PDZ domain / SH3 domain / PDZ domain profile. / Domain present in PSD-95, Dlg, and ZO-1/2. / PDZ domain / PDZ superfamily / Src homology 3 domains / SH3-like domain superfamily / Src homology 3 (SH3) domain profile. / SH3 domain / Roll / P-loop containing nucleoside triphosphate hydrolase / Mainly BetaSimilarity search - Domain/homology |
|---|
| Biological species |   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat)
synthetic construct (others) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 2.151 Å molecular replacement / Resolution: 2.151 Å |
|---|
 Authors Authors | De, S. / Spaller, M.R. / Olson, R. |
|---|
 Citation Citation |  Journal: to be published Journal: to be published
Title: Chemically-modified peptides as inhibitors of PDZ3 of PSD-95
Authors: Memic, A. / De, S. / Lakhani, B. / Beveridge, D.L. / Olson, R. / Spaller, M.R. |
|---|
| History | | Deposition | Aug 3, 2015 | Deposition site: RCSB / Processing site: RCSB |
|---|
| Revision 1.0 | Aug 17, 2016 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Sep 27, 2023 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Refinement description
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / diffrn_radiation_wavelength / pdbx_initial_refinement_model / pdbx_struct_oper_list / struct_conn
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_oper_list.symmetry_operation / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_seq_id |
|---|
| Revision 2.0 | Apr 24, 2024 | Group: Advisory / Atomic model ...Advisory / Atomic model / Data collection / Database references / Derived calculations / Other / Polymer sequence / Refinement description / Source and taxonomy / Structure summary
Category: atom_site / atom_site_anisotrop ...atom_site / atom_site_anisotrop / cell / entity / entity_poly / entity_src_gen / pdbx_entity_nonpoly / pdbx_nonpoly_scheme / pdbx_poly_seq_scheme / pdbx_refine_tls / pdbx_refine_tls_group / pdbx_struct_assembly / pdbx_struct_assembly_gen / pdbx_struct_assembly_prop / pdbx_unobs_or_zero_occ_atoms / pdbx_unobs_or_zero_occ_residues / struct_asym / struct_conn / struct_ref_seq
Item: _atom_site.auth_asym_id / _atom_site.group_PDB ..._atom_site.auth_asym_id / _atom_site.group_PDB / _atom_site.label_asym_id / _atom_site.label_entity_id / _atom_site.label_seq_id / _atom_site_anisotrop.pdbx_auth_asym_id / _atom_site_anisotrop.pdbx_label_asym_id / _atom_site_anisotrop.pdbx_label_seq_id / _cell.Z_PDB / _entity_poly.pdbx_strand_id / _entity_src_gen.gene_src_common_name / _pdbx_refine_tls.origin_x / _pdbx_refine_tls.origin_y / _pdbx_refine_tls.origin_z / _pdbx_refine_tls_group.beg_auth_seq_id / _pdbx_refine_tls_group.end_auth_seq_id / _pdbx_refine_tls_group.selection_details / _pdbx_struct_assembly.oligomeric_count / _pdbx_struct_assembly.oligomeric_details / _pdbx_struct_assembly_gen.asym_id_list / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_seq_id |
|---|
| Revision 3.0 | Jul 10, 2024 | Group: Data collection / Non-polymer description / Structure summary
Category: chem_comp / chem_comp_atom ...chem_comp / chem_comp_atom / chem_comp_bond / entity
Item: _chem_comp.formula / _chem_comp.formula_weight / _entity.formula_weight |
|---|
| Revision 3.1 | Oct 16, 2024 | Group: Structure summary / Category: pdbx_entry_details / pdbx_modification_feature |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT /
MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 2.151 Å
molecular replacement / Resolution: 2.151 Å  Authors
Authors Citation
Citation Journal: to be published
Journal: to be published Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5d13.cif.gz
5d13.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5d13.ent.gz
pdb5d13.ent.gz PDB format
PDB format 5d13.json.gz
5d13.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/d1/5d13
https://data.pdbj.org/pub/pdb/validation_reports/d1/5d13 ftp://data.pdbj.org/pub/pdb/validation_reports/d1/5d13
ftp://data.pdbj.org/pub/pdb/validation_reports/d1/5d13
 Links
Links Assembly
Assembly




 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  NSLS
NSLS  / Beamline: X25 / Wavelength: 1.1 Å
/ Beamline: X25 / Wavelength: 1.1 Å molecular replacement
molecular replacement Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller










 PDBj
PDBj