+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4cse | ||||||
|---|---|---|---|---|---|---|---|
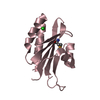
| Title | PIH N-terminal domain | ||||||
 Components Components |
| ||||||
 Keywords Keywords | CHAPERONE / MOLECULAR CHAPERONES / MULTIPROTEIN COMPLEXES / PHOSPHORYLATION | ||||||
| Function / homology |  Function and homology information Function and homology informationprotein-containing complex stabilizing activity / TORC1 complex assembly / snoRNA localization / 'de novo' cotranslational protein folding / positive regulation of glucose mediated signaling pathway / TTT Hsp90 cochaperone complex / pre-snoRNP complex / R2TP complex / positive regulation of transcription of nucleolar large rRNA by RNA polymerase I / RPAP3/R2TP/prefoldin-like complex ...protein-containing complex stabilizing activity / TORC1 complex assembly / snoRNA localization / 'de novo' cotranslational protein folding / positive regulation of glucose mediated signaling pathway / TTT Hsp90 cochaperone complex / pre-snoRNP complex / R2TP complex / positive regulation of transcription of nucleolar large rRNA by RNA polymerase I / RPAP3/R2TP/prefoldin-like complex / box C/D snoRNP assembly / telomeric repeat DNA binding / histone reader activity / epithelial cell differentiation / positive regulation of TORC1 signaling / phosphoprotein binding / Hsp90 protein binding / kinase binding / rRNA processing / ATPase binding / histone binding / molecular adaptor activity / chromosome, telomeric region / protein stabilization / nuclear body / chromatin remodeling / ribonucleoprotein complex / protein kinase binding / protein-containing complex binding / nucleolus / membrane / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3.3 Å MOLECULAR REPLACEMENT / Resolution: 3.3 Å | ||||||
 Authors Authors | Morgan, R.M. / Roe, S.M. | ||||||
 Citation Citation |  Journal: Structure / Year: 2014 Journal: Structure / Year: 2014Title: Structural Basis for Phosphorylation-Dependent Recruitment of Tel2 to Hsp90 by Pih1. Authors: Pal, M. / Morgan, M. / Phelps, S.E. / Roe, S.M. / Parry-Morris, S. / Downs, J.A. / Polier, S. / Pearl, L.H. / Prodromou, C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4cse.cif.gz 4cse.cif.gz | 58.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4cse.ent.gz pdb4cse.ent.gz | 41.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4cse.json.gz 4cse.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/cs/4cse https://data.pdbj.org/pub/pdb/validation_reports/cs/4cse ftp://data.pdbj.org/pub/pdb/validation_reports/cs/4cse ftp://data.pdbj.org/pub/pdb/validation_reports/cs/4cse | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4cguC  4cgvC  4cgwC  4chhC  4cktSC  4cv4C C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14976.923 Da / Num. of mol.: 2 / Fragment: RESIDUES 47-179 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Protein/peptide | Mass: 1135.972 Da / Num. of mol.: 2 / Fragment: RESIDUES 498-506 / Source method: obtained synthetically / Source: (synth.)  #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.76 Å3/Da / Density % sol: 55 % / Description: NONE |
|---|---|
| Crystal grow | pH: 7 Details: 0.15M SODIUM POTASSIUM PHOSPHATE, 20% PEG 3350, 0.1M BIS-TRIS PH 6.5 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I04-1 / Wavelength: 0.92 / Beamline: I04-1 / Wavelength: 0.92 |
| Detector | Type: DECTRIS PILATUS 2M / Detector: PIXEL / Date: Feb 5, 2013 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.92 Å / Relative weight: 1 |
| Reflection | Resolution: 3.3→56.76 Å / Num. obs: 5693 / % possible obs: 99.6 % / Observed criterion σ(I): 2 / Redundancy: 11.7 % / Biso Wilson estimate: 66.3 Å2 / Rmerge(I) obs: 0.17 / Net I/σ(I): 12.8 |
| Reflection shell | Resolution: 3.3→3.48 Å / Redundancy: 12.5 % / Rmerge(I) obs: 0.6 / Mean I/σ(I) obs: 4.7 / % possible all: 99.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 4CKT Resolution: 3.3→56.759 Å / SU ML: 0.48 / σ(F): 1.36 / Phase error: 30.68 / Stereochemistry target values: ML
| ||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3.3→56.759 Å
| ||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

