[English] 日本語
 Yorodumi
Yorodumi- PDB-3e85: Crystal Structure of Pathogenesis-related Protein LlPR-10.2B from... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3.0E+85 | ||||||
|---|---|---|---|---|---|---|---|
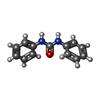
| Title | Crystal Structure of Pathogenesis-related Protein LlPR-10.2B from yellow lupine in complex with Diphenylurea | ||||||
 Components Components | PR10.2B | ||||||
 Keywords Keywords | PLANT PROTEIN / PLANT HORMONES / CYTOKININ / DIPHENYLUREA / PLANT PR-10 PROTEIN / YELLOW LUPINE / Pathogenesis-related protein / Plant defense | ||||||
| Function / homology |  Function and homology information Function and homology informationcytokinin binding / melatonin binding / abscisic acid binding / Hydrolases; Acting on ester bonds; Endoribonucleases producing 3'-phosphomonoesters / abscisic acid-activated signaling pathway / protein phosphatase inhibitor activity / RNA nuclease activity / defense response / signaling receptor activity / calcium ion binding ...cytokinin binding / melatonin binding / abscisic acid binding / Hydrolases; Acting on ester bonds; Endoribonucleases producing 3'-phosphomonoesters / abscisic acid-activated signaling pathway / protein phosphatase inhibitor activity / RNA nuclease activity / defense response / signaling receptor activity / calcium ion binding / nucleus / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.95 Å MOLECULAR REPLACEMENT / Resolution: 1.95 Å | ||||||
 Authors Authors | Fernandes, H.C. / Bujacz, G. / Bujacz, A. / Sikorski, M.M. / Jaskolski, M. | ||||||
 Citation Citation |  Journal: Febs J. / Year: 2009 Journal: Febs J. / Year: 2009Title: Cytokinin-induced structural adaptability of a Lupinus luteus PR-10 protein. Authors: Fernandes, H. / Bujacz, A. / Bujacz, G. / Jelen, F. / Jasinski, M. / Kachlicki, P. / Otlewski, J. / Sikorski, M.M. / Jaskolski, M. #1:  Journal: J.Mol.Biol. / Year: 2008 Journal: J.Mol.Biol. / Year: 2008Title: Lupinus luteus Pathogenesis-Related Protein as a Reservoir for Cytokinin Authors: Fernandes, H. / Pasternak, O. / Bujacz, G. / Bujacz, A. / Sikorski, M.M. / Jaskolski, M. #2:  Journal: Acta Crystallogr.,Sect.D / Year: 2005 Journal: Acta Crystallogr.,Sect.D / Year: 2005Title: Structure of a yellow lupin pathogenesis-related PR-10 protein belonging to a novel subclass Authors: Pasternak, O. / Biesiadka, J. / Dolot, R. / Handschuh, L. / Bujacz, G. / Sikorski, M.M. / Jaskolski, M. #3:  Journal: J.Mol.Biol. / Year: 2002 Journal: J.Mol.Biol. / Year: 2002Title: Crystal structures of two homologous pathogenesis-related proteins from yellow lupine Authors: Biesiadka, J. / Bujacz, G. / Sikorski, M.M. / Jaskolski, M. #4:  Journal: Plant Cell / Year: 2006 Journal: Plant Cell / Year: 2006Title: Crystal structure of Vigna radiata cytokinin-specific binding protein in complex with zeatin Authors: Pasternak, O. / Bujacz, G.D. / Fujimoto, Y. / Hashimoto, Y. / Jelen, F. / Otlewski, J. / Sikorski, M.M. / Jaskolski, M. #5:  Journal: Nat.Struct.Mol.Biol. / Year: 1996 Journal: Nat.Struct.Mol.Biol. / Year: 1996Title: X-ray and NMR structure of Bet v 1, the origin of birch pollen allergy Authors: Gajhede, M. / Osmark, P. / Poulsen, F.M. / Ipsen, H. / Larsen, J.N. / Joost van Neerven, R.J. / Schou, C. / Lowenstein, H. / Spangfort, M.D. #6:  Journal: J.Mol.Biol. / Year: 2003 Journal: J.Mol.Biol. / Year: 2003Title: Crystal structure of a hypoallergenic isoform of the major birch pollen allergen Bet v 1 and its likely biological function as a plant steroid carrier Authors: Markovi-Housley, Z. / Degano, M. / Lamba, D. / von Roepenack-Lahaye, E. / Clemens, S. / Susani, M. / Ferreira, F. / Scheiner, O. / Breiteneder, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3e85.cif.gz 3e85.cif.gz | 50.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3e85.ent.gz pdb3e85.ent.gz | 35.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3e85.json.gz 3e85.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  3e85_validation.pdf.gz 3e85_validation.pdf.gz | 448.6 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  3e85_full_validation.pdf.gz 3e85_full_validation.pdf.gz | 451.7 KB | Display | |
| Data in XML |  3e85_validation.xml.gz 3e85_validation.xml.gz | 10.5 KB | Display | |
| Data in CIF |  3e85_validation.cif.gz 3e85_validation.cif.gz | 14.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/e8/3e85 https://data.pdbj.org/pub/pdb/validation_reports/e8/3e85 ftp://data.pdbj.org/pub/pdb/validation_reports/e8/3e85 ftp://data.pdbj.org/pub/pdb/validation_reports/e8/3e85 | HTTPS FTP |
-Related structure data
| Related structure data |  2qimS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 16906.000 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-BSU / #3: Chemical | #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.86 Å3/Da / Density % sol: 33.88 % |
|---|---|
| Crystal grow | Temperature: 292 K / Method: vapor diffusion, hanging drop / pH: 6 Details: 1.4 M Sodium citrate, 0.1 M citrate buffer, pH 6.0, VAPOR DIFFUSION, HANGING DROP, temperature 292K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: X11 / Wavelength: 0.81 Å / Beamline: X11 / Wavelength: 0.81 Å |
| Detector | Type: MAR CCD 165 mm / Detector: CCD / Date: Oct 16, 2005 / Details: mirrors |
| Radiation | Monochromator: Si [111], horizontally focussing / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.81 Å / Relative weight: 1 |
| Reflection | Resolution: 1.95→40 Å / Num. obs: 9529 / % possible obs: 99.56 % / Observed criterion σ(I): -3 / Redundancy: 5.7 % / Biso Wilson estimate: 37.9 Å2 / Rmerge(I) obs: 0.075 / Net I/σ(I): 20.8 |
| Reflection shell | Resolution: 1.95→2.02 Å / Redundancy: 5.4 % / Rmerge(I) obs: 0.398 / Mean I/σ(I) obs: 3.8 / % possible all: 96.8 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 2QIM Resolution: 1.95→15 Å / Cor.coef. Fo:Fc: 0.958 / Cor.coef. Fo:Fc free: 0.932 / SU B: 9.931 / SU ML: 0.151 / TLS residual ADP flag: LIKELY RESIDUAL / Cross valid method: THROUGHOUT / ESU R: 0.249 / ESU R Free: 0.198 / Stereochemistry target values: Engh & Huber / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 30.611 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.95→15 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.95→2 Å / Total num. of bins used: 20
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller
















 PDBj
PDBj