[English] 日本語
 Yorodumi
Yorodumi- PDB-2xum: FACTOR INHIBITING HIF (FIH) Q239H MUTANT IN COMPLEX WITH ZN(II), ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2xum | ||||||
|---|---|---|---|---|---|---|---|
| Title | FACTOR INHIBITING HIF (FIH) Q239H MUTANT IN COMPLEX WITH ZN(II), NOG AND ASP-SUBSTRATE PEPTIDE (20-MER) | ||||||
 Components Components |
| ||||||
 Keywords Keywords | OXIDOREDUCTASE/PEPTIDE / OXIDOREDUCTASE-PEPTIDE COMPLEX / NON-HEME / DIOXYGENASE / OXYGENASE / HYPOXIA / METAL-BINDING / HELIX-LOOP-HELIX-BETA / FACIAL TRIAD SIGNALING / BETA-HYDROXYLATION / TRANSCRIPTION ACTIVATOR/INHIBITOR | ||||||
| Function / homology |  Function and homology information Function and homology informationhypoxia-inducible factor-asparagine dioxygenase / : / [protein]-asparagine 3-dioxygenase activity / peptidyl-histidine dioxygenase activity / peptidyl-aspartic acid 3-dioxygenase activity / regulation of vascular endothelial growth factor receptor signaling pathway / Cellular response to hypoxia / positive regulation of vasculogenesis / carboxylic acid binding / ankyrin repeat binding ...hypoxia-inducible factor-asparagine dioxygenase / : / [protein]-asparagine 3-dioxygenase activity / peptidyl-histidine dioxygenase activity / peptidyl-aspartic acid 3-dioxygenase activity / regulation of vascular endothelial growth factor receptor signaling pathway / Cellular response to hypoxia / positive regulation of vasculogenesis / carboxylic acid binding / ankyrin repeat binding / Notch binding / oxygen sensor activity / negative regulation of Notch signaling pathway / positive regulation of myoblast differentiation / NF-kappaB binding / ferrous iron binding / transcription corepressor activity / RNA polymerase II-specific DNA-binding transcription factor binding / perinuclear region of cytoplasm / negative regulation of transcription by RNA polymerase II / protein homodimerization activity / zinc ion binding / nucleoplasm / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human)SYNTHETIC CONSTRUCT (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Chowdhury, R. / McDonough, M.A. / Schofield, C.J. | ||||||
 Citation Citation |  Journal: J. Biol. Chem. / Year: 2011 Journal: J. Biol. Chem. / Year: 2011Title: Asparagine and aspartate hydroxylation of the cytoskeletal ankyrin family is catalyzed by factor-inhibiting hypoxia-inducible factor. Authors: Yang, M. / Ge, W. / Chowdhury, R. / Claridge, T.D. / Kramer, H.B. / Schmierer, B. / McDonough, M.A. / Gong, L. / Kessler, B.M. / Ratcliffe, P.J. / Coleman, M.L. / Schofield, C.J. #1:  Journal: J.Biol.Chem. / Year: 2007 Journal: J.Biol.Chem. / Year: 2007Title: Asparaginyl Hydroxylation of the Notch Ankyrin Repeat Domain by Factor Inhibiting Hypoxia-Inducible Factor. Authors: Coleman, M.L. / McDonough, M.A. / Hewitson, K.S. / Coles, C. / Mecinovic, J. / Edelmann, M. / Cook, K.M. / Cockman, M.E. / Lancaster, D.E. / Kessler, B.M. / Oldham, N.J. / Ratcliffe, P.J. / Schofield, C.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2xum.cif.gz 2xum.cif.gz | 92 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2xum.ent.gz pdb2xum.ent.gz | 68.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2xum.json.gz 2xum.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xu/2xum https://data.pdbj.org/pub/pdb/validation_reports/xu/2xum ftp://data.pdbj.org/pub/pdb/validation_reports/xu/2xum ftp://data.pdbj.org/pub/pdb/validation_reports/xu/2xum | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1h2kS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein / Protein/peptide , 2 types, 2 molecules AS
| #1: Protein | Mass: 40338.297 Da / Num. of mol.: 1 / Mutation: YES Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  References: UniProt: Q9NWT6, peptide-aspartate beta-dioxygenase |
|---|---|
| #2: Protein/peptide | Mass: 2233.519 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: BASED ON BIOLOGICAL SEQUENCE / Source: (synth.) SYNTHETIC CONSTRUCT (others) |
-Non-polymers , 4 types, 159 molecules 
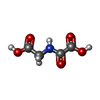





| #3: Chemical | ChemComp-ZN / | ||
|---|---|---|---|
| #4: Chemical | ChemComp-OGA / | ||
| #5: Chemical | | #6: Water | ChemComp-HOH / | |
-Details
| Compound details | ENGINEERED| Nonpolymer details | RESIDUE OGA IS THE N-OXALYL GLYCINE (NOG). | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.22 Å3/Da / Density % sol: 62 % / Description: NONE |
|---|---|
| Crystal grow | Method: vapor diffusion, sitting drop / pH: 7.5 Details: 1.6M AMMONIUM SULPHATE, 6% PEG400, 0.1M HEPES PH7.5, VAPOR DIFFUSION, SITTING DROP |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I04 / Wavelength: 0.9795 / Beamline: I04 / Wavelength: 0.9795 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Sep 25, 2010 / Details: MIRRORS |
| Radiation | Monochromator: SI 111 / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9795 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→74.12 Å / Num. obs: 26863 / % possible obs: 99.1 % / Observed criterion σ(I): 2 / Redundancy: 9.2 % / Biso Wilson estimate: 51.7 Å2 / Rmerge(I) obs: 0.07 / Net I/σ(I): 28.5 |
| Reflection shell | Resolution: 2.2→2.28 Å / Redundancy: 8.4 % / Rmerge(I) obs: 0.88 / Mean I/σ(I) obs: 2.23 / % possible all: 98 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1H2K Resolution: 2.2→74.12 Å / Cor.coef. Fo:Fc: 0.957 / Cor.coef. Fo:Fc free: 0.948 / SU B: 11.189 / SU ML: 0.151 / Cross valid method: THROUGHOUT / ESU R: 0.215 / ESU R Free: 0.172 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 49.924 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→74.12 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller
























 PDBj
PDBj



