| Entry | Database: PDB / ID: 2vn6
|
|---|
| Title | The Clostridium cellulolyticum dockerin displays a dual binding mode for its cohesin partner |
|---|
 Components Components | - ENDOGLUCANASE A
- SCAFFOLDING PROTEIN
|
|---|
 Keywords Keywords | CELL ADHESION / CARBOHYDRATE METABOLISM / POLYSACCHARIDE DEGRADATION / HYDROLASE / GLYCOSIDASE / CELLULOSE DEGRADATION |
|---|
| Function / homology |  Function and homology information Function and homology information
cellulose binding / cellulase / cellulase activity / beta-glucosidase activity / cellulose catabolic process / cell surface / extracellular region / metal ion bindingSimilarity search - Function Type 1 dockerin domain / Dockerin domain / Carbohydrate binding X2 domain / Carbohydrate binding domain X2 / Immunoglobulin-like - #680 / Cellulosome anchoring protein, cohesin domain / Cohesin domain / Cellulose binding domain / Carbohydrate-binding module 3 / Cellulose binding domain ...Type 1 dockerin domain / Dockerin domain / Carbohydrate binding X2 domain / Carbohydrate binding domain X2 / Immunoglobulin-like - #680 / Cellulosome anchoring protein, cohesin domain / Cohesin domain / Cellulose binding domain / Carbohydrate-binding module 3 / Cellulose binding domain / CBM3 (carbohydrate binding type-3) domain profile. / Carbohydrate-binding module 3 superfamily / : / Clostridium cellulosome enzymes repeated domain signature. / Glycoside hydrolase, family 5, conserved site / Glycosyl hydrolases family 5 signature. / Dockerin domain / Dockerin domain profile. / Dockerin type I domain / Dockerin type I repeat / Dockerin domain superfamily / CBM2/CBM3, carbohydrate-binding domain superfamily / Cellulase (glycosyl hydrolase family 5) / Glycoside hydrolase, family 5 / EF-Hand 1, calcium-binding site / EF-hand calcium-binding domain. / Immunoglobulin E-set / Glycoside hydrolase superfamily / Immunoglobulin-like fold / Immunoglobulin-like / Sandwich / Orthogonal Bundle / Mainly Beta / Mainly AlphaSimilarity search - Domain/homology |
|---|
| Biological species |  CLOSTRIDIUM CELLULOLYTICUM (bacteria) CLOSTRIDIUM CELLULOLYTICUM (bacteria) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.49 Å MOLECULAR REPLACEMENT / Resolution: 1.49 Å |
|---|
 Authors Authors | Pinheiro, B.A. / Prates, J.A.M. / Proctor, M.R. / Gilbert, H.J. / Davies, G.J. / Money, V.A. / Martinez-Fleites, C. / Bayer, E.A. / Fontes, C.M.G.A. / Fierobe, H.P. |
|---|
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2008 Journal: J.Biol.Chem. / Year: 2008
Title: The Clostridium Cellulolyticum Dockerin Displays a Dual Binding Mode for its Cohesin Partner.
Authors: Pinheiro, B.A. / Proctor, M.R. / Martinez-Fleites, C. / Prates, J.A.M. / Money, V.A. / Davies, G.J. / Bayer, E.A. / Fontes, C.M.G.A. / Fierobe, H.P. / Gilbert, H.J. |
|---|
| History | | Deposition | Jan 31, 2008 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | May 13, 2008 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | May 8, 2011 | Group: Version format compliance |
|---|
| Revision 1.2 | Jul 13, 2011 | Group: Version format compliance |
|---|
| Revision 1.3 | May 8, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Other
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_struct_conn_angle / struct_conn
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information CLOSTRIDIUM CELLULOLYTICUM (bacteria)
CLOSTRIDIUM CELLULOLYTICUM (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.49 Å
MOLECULAR REPLACEMENT / Resolution: 1.49 Å  Authors
Authors Citation
Citation Journal: J.Biol.Chem. / Year: 2008
Journal: J.Biol.Chem. / Year: 2008 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2vn6.cif.gz
2vn6.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2vn6.ent.gz
pdb2vn6.ent.gz PDB format
PDB format 2vn6.json.gz
2vn6.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/vn/2vn6
https://data.pdbj.org/pub/pdb/validation_reports/vn/2vn6 ftp://data.pdbj.org/pub/pdb/validation_reports/vn/2vn6
ftp://data.pdbj.org/pub/pdb/validation_reports/vn/2vn6 Links
Links Assembly
Assembly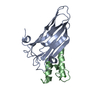
 Components
Components CLOSTRIDIUM CELLULOLYTICUM (bacteria) / Production host:
CLOSTRIDIUM CELLULOLYTICUM (bacteria) / Production host: 
 CLOSTRIDIUM CELLULOLYTICUM (bacteria) / Production host:
CLOSTRIDIUM CELLULOLYTICUM (bacteria) / Production host: 
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  ESRF
ESRF  / Beamline: ID14-2 / Wavelength: 0.933
/ Beamline: ID14-2 / Wavelength: 0.933  Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller
















 PDBj
PDBj


