+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2pkv | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | D-(GGTATACC) ambient pressure | ||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / HIGH PRESSURE | Function / homology | DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  FOURIER SYNTHESIS / Resolution: 1.6 Å FOURIER SYNTHESIS / Resolution: 1.6 Å  Authors AuthorsGirard, E. / Prange, T. / Kahn, R. / Fourme, R. |  Citation Citation Journal: Nucleic Acids Res. / Year: 2007 Journal: Nucleic Acids Res. / Year: 2007Title: Adaptation of the base-paired double-helix molecular architecture to extreme pressure. Authors: Girard, E. / Prange, T. / Dhaussy, A.C. / Migianu-Griffoni, E. / Lecouvey, M. / Chervin, J.C. / Mezouar, M. / Kahn, R. / Fourme, R. #1: Journal: Nature / Year: 1989 Title: Coexistence of A-and B-Form DNA in a Single Crystal Lattice Authors: Doucet, J. / Benoit, J.-P. / Cruse, W.B.T. / Prange, T. / Kennard, O. #2: Journal: J.Synchrotron Radia. / Year: 2001 Title: High-pressure protein crystallography (HPPX): Instrumentation, methodology and results on lysozyme crystals Authors: Fourme, R. / Kahn, R. / Mezouar, M. / Girard, E. / Horentrup, C. / Prange, T. / Ascone, I. #3:  Journal: BIOCHEM.BIOPHYS.ACTA PROTEINS & PROTEOMICS / Year: 2006 Journal: BIOCHEM.BIOPHYS.ACTA PROTEINS & PROTEOMICS / Year: 2006Title: High pressure macromolecular crystallography: The 140 MPa crystal structure at 2.3 A resolution of urate oxidase, A 135 KD tetrameric assembly Authors: Colloc'h, N. / Girard, E. / Dhaussy, A.C. / Kahn, R. / Ascone, I. / Mezouar, M. / Fourme, R. #4:  Journal: J.Mol.Biol. / Year: 1987 Journal: J.Mol.Biol. / Year: 1987Title: Crystal structure of hen egg-white lysozyme at a hydrostatic pressure of 1000 atmospheres. Authors: Kundrot, C.E. / Richards, F.M. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2pkv.cif.gz 2pkv.cif.gz | 19.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2pkv.ent.gz pdb2pkv.ent.gz | 12.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2pkv.json.gz 2pkv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2pkv_validation.pdf.gz 2pkv_validation.pdf.gz | 367.3 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2pkv_full_validation.pdf.gz 2pkv_full_validation.pdf.gz | 370.8 KB | Display | |
| Data in XML |  2pkv_validation.xml.gz 2pkv_validation.xml.gz | 3.6 KB | Display | |
| Data in CIF |  2pkv_validation.cif.gz 2pkv_validation.cif.gz | 4.4 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pk/2pkv https://data.pdbj.org/pub/pdb/validation_reports/pk/2pkv ftp://data.pdbj.org/pub/pdb/validation_reports/pk/2pkv ftp://data.pdbj.org/pub/pdb/validation_reports/pk/2pkv | HTTPS FTP |
-Related structure data
| Related structure data |  2pl4C  2pl8C  2plbC 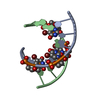 115dS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Details | THE BIOLOGICAL ASSEMBLY IS A DIMER |
- Components
Components
| #1: DNA chain | Mass: 2426.617 Da / Num. of mol.: 2 / Source method: obtained synthetically / Details: SYNTHETIC DNA #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.42 Å3/Da / Density % sol: 49.21 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 290 K / Method: vapor diffusion, sitting drop / pH: 7 Details: 10 mg of lyophilized DNA dissolved in 0.2 ml of a 15 % MPD solution with additives: 10-5 M sodium azide, 10-2 M MgCl2 and 2.10-2 spermine chloride. Reservoir : same solution but 50 % MPD, pH ...Details: 10 mg of lyophilized DNA dissolved in 0.2 ml of a 15 % MPD solution with additives: 10-5 M sodium azide, 10-2 M MgCl2 and 2.10-2 spermine chloride. Reservoir : same solution but 50 % MPD, pH 7, VAPOR DIFFUSION, SITTING DROP, temperature 290K | ||||||||||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 295 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID27 / Wavelength: 0.3738 Å / Beamline: ID27 / Wavelength: 0.3738 Å |
| Detector | Type: MAR scanner 345 mm plate / Detector: IMAGE PLATE / Date: Nov 10, 2006 |
| Radiation | Monochromator: SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.3738 Å / Relative weight: 1 |
| Reflection | Resolution: 1.6→19 Å / Num. all: 6228 / Num. obs: 6228 / % possible obs: 94.5 % / Observed criterion σ(F): 2 / Observed criterion σ(I): 4 / Redundancy: 4 % / Rmerge(I) obs: 0.045 / Net I/σ(I): 9.3 |
| Reflection shell | Resolution: 1.6→1.7 Å / Redundancy: 4 % / Rmerge(I) obs: 0.33 / Mean I/σ(I) obs: 2.3 / % possible all: 96.2 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  FOURIER SYNTHESIS FOURIER SYNTHESISStarting model: 115D FROM PDB Resolution: 1.6→10 Å / Num. parameters: 1523 / Num. restraintsaints: 1585 / Isotropic thermal model: individual B factors / Cross valid method: FREE R / σ(F): 2 / σ(I): 4 / Stereochemistry target values: ENGH AND HUBER
| |||||||||||||||||||||||||||||||||
| Refine analyze | Num. disordered residues: 0 / Occupancy sum hydrogen: 184 / Occupancy sum non hydrogen: 379 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.6→10 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller







 PDBj
PDBj


