+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2org | ||||||
|---|---|---|---|---|---|---|---|
| Title | Directing Macromolecular Conformation Through Halogen Bonds | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA / DNA Holliday Junction / Halogen Bond | ||||||
| Function / homology | DNA Function and homology information Function and homology information | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Voth, A.R. / Hays, F.A. / Ho, P.S. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.Usa / Year: 2007 Journal: Proc.Natl.Acad.Sci.Usa / Year: 2007Title: Directing macromolecular conformation through halogen bonds. Authors: Voth, A.R. / Hays, F.A. / Ho, P.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2org.cif.gz 2org.cif.gz | 22.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2org.ent.gz pdb2org.ent.gz | 13.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2org.json.gz 2org.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/or/2org https://data.pdbj.org/pub/pdb/validation_reports/or/2org ftp://data.pdbj.org/pub/pdb/validation_reports/or/2org ftp://data.pdbj.org/pub/pdb/validation_reports/or/2org | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2orfC  2orhC  1p54S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 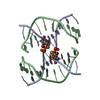
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
| ||||||||
| Details | The second half of the biological assembly (the Holliday junction) is generated by the two fold rotation: -x, y, -z+1 |
- Components
Components
| #1: DNA chain | Mass: 3125.874 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: Synthetically prepared decanucleotide sequences. |
|---|---|
| #2: DNA chain | Mass: 3029.994 Da / Num. of mol.: 1 / Source method: obtained synthetically |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.17 Å3/Da / Density % sol: 43.42 % | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 7 Details: 25 mM sodium cacodylate, 20 mM calcium chloride, 1.2 mM spermine, and 0.35 mM of each DNA strand., pH 7.0, VAPOR DIFFUSION, SITTING DROP, temperature 298K | ||||||||||||||||||||||||||||
| Components of the solutions |
|
-Data collection
| Diffraction | Mean temperature: 133 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å ROTATING ANODE / Type: RIGAKU / Wavelength: 1.5418 Å |
| Detector | Type: RIGAKU RAXIS IV / Detector: IMAGE PLATE / Date: Jul 13, 2004 / Details: Osmic mirrors |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2→18.2 Å / Num. all: 3852 / Num. obs: 3362 / % possible obs: 87.3 % / Redundancy: 2.4 % / Biso Wilson estimate: 5.9 Å2 / Rmerge(I) obs: 0.049 |
| Reflection shell | Resolution: 2→2.07 Å / Redundancy: 1.8 % / Rmerge(I) obs: 0.281 / Mean I/σ(I) obs: 2.3 / Num. unique all: 382 / % possible all: 49.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1P54 Resolution: 2→18.2 Å / Rfactor Rfree error: 0.015 / Data cutoff high absF: 43547.84 / Data cutoff high rms absF: 43547.84 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 94.7022 Å2 / ksol: 0.357361 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 14.3 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→18.2 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.09 Å / Rfactor Rfree error: 0.054 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller








 PDBj
PDBj


