+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2fmb | ||||||
|---|---|---|---|---|---|---|---|
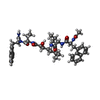
| Title | EIAV PROTEASE COMPLEXED WITH AN INHIBITOR LP-130 | ||||||
 Components Components | EQUINE INFECTIOUS ANEMIA VIRUS PROTEASE | ||||||
 Keywords Keywords | ASPARTIC PROTEASE / RETROPEPSIN / RETROVIRUS / EIAV / HORSE | ||||||
| Function / homology |  Function and homology information Function and homology informationHydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / RNA-directed DNA polymerase activity ...Hydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / DNA recombination / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / symbiont entry into host cell / proteolysis / DNA binding / zinc ion binding Similarity search - Function | ||||||
| Biological species |  Equine infectious anemia virus Equine infectious anemia virus | ||||||
| Method |  X-RAY DIFFRACTION / RIGID BODY REFINEMENT / Resolution: 1.8 Å X-RAY DIFFRACTION / RIGID BODY REFINEMENT / Resolution: 1.8 Å | ||||||
 Authors Authors | Kervinen, J. / Lubkowski, J. / Zdanov, A. / Wlodawer, A. / Gustchina, A. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 1998 Journal: Protein Sci. / Year: 1998Title: Toward a universal inhibitor of retroviral proteases: comparative analysis of the interactions of LP-130 complexed with proteases from HIV-1, FIV, and EIAV. Authors: Kervinen, J. / Lubkowski, J. / Zdanov, A. / Bhatt, D. / Dunn, B.M. / Hui, K.Y. / Powell, D.J. / Kay, J. / Wlodawer, A. / Gustchina, A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2fmb.cif.gz 2fmb.cif.gz | 38.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2fmb.ent.gz pdb2fmb.ent.gz | 25.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2fmb.json.gz 2fmb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/fm/2fmb https://data.pdbj.org/pub/pdb/validation_reports/fm/2fmb ftp://data.pdbj.org/pub/pdb/validation_reports/fm/2fmb ftp://data.pdbj.org/pub/pdb/validation_reports/fm/2fmb | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1odyC  4fivC  1fmbS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
| ||||||||
| Components on special symmetry positions |
| ||||||||
| Details | THIS FILE CONTAINS ONLY A MONOMER, WHILE THE ACTIVE ENZYME IS A DIMER. IN ORDER TO CREATE A DIMERIC MOLECULE, CRYSTALLOGRAPHIC COORDINATES NEED TO BE TRANSFORMED TO (-X), Y, (-Z). THE TWO ORIENTATIONS OF THE INHIBITOR CREATED IN THAT MANNER OVERLAP, WITH COMPLETE SUPERPOSITION AT THE PERIPHERY AND SOME DEVIATION IN THE CENTER. RESIDUES 56 - 58 HAVE BEEN MODELED WITH TWO ORIENTATIONS OF THEIR MAIN CHAINS, SINCE THEY FORM ALTERNATE HYDROGEN BONDS, DUE TO THE SAME DISORDER PHENOMENON. |
- Components
Components
| #1: Protein | Mass: 11392.265 Da / Num. of mol.: 1 / Mutation: I54G Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Equine infectious anemia virus / Genus: Lentivirus / Production host: Equine infectious anemia virus / Genus: Lentivirus / Production host:  References: UniProt: P32542, UniProt: Q66729*PLUS, HIV-1 retropepsin |
|---|---|
| #2: Chemical | ChemComp-LP1 / |
| #3: Water | ChemComp-HOH / |
| Nonpolymer details | KI VALUE OF LP-130 FOR EIAV PR IS 2 NM |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.26 Å3/Da / Density % sol: 45.54 % | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 4.6 / Details: pH 4.6 | ||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ENRAF-NONIUS FR591 / Wavelength: 1.5418 ROTATING ANODE / Type: ENRAF-NONIUS FR591 / Wavelength: 1.5418 |
| Detector | Type: MACSCIENCE / Detector: IMAGE PLATE / Date: Sep 18, 1996 / Details: MIRRORS |
| Radiation | Monochromator: NI FILTER / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→20 Å / Num. obs: 8666 / % possible obs: 90.9 % / Observed criterion σ(I): 1 / Rmerge(I) obs: 0.049 |
| Reflection shell | Resolution: 1.8→1.83 Å / Rmerge(I) obs: 0.3 / % possible all: 81.4 |
| Reflection shell | *PLUS % possible obs: 91.4 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: RIGID BODY REFINEMENT Starting model: PDB ENTRY 1FMB Resolution: 1.8→10 Å / σ(F): 3
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→10 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: PROFFT / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.143 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller













 PDBj
PDBj