Entry Database : PDB / ID : 2d2cTitle Crystal Structure Of Cytochrome B6F Complex with DBMIB From M. Laminosus (Cytochrome b6-f complex subunit ...) x 5 Apocytochrome f Cytochrome b6 Cytochrome b6-f complex iron-sulfur subunit Keywords Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Mastigocladus laminosus (bacteria)Method / / Resolution : 3.8 Å Authors Yan, J. / Kurisu, G. / Cramer, W.A. Journal : Proc.Natl.Acad.Sci.USA / Year : 2006Title : Intraprotein transfer of the quinone analogue inhibitor 2,5-dibromo-3-methyl-6-isopropyl-p-benzoquinone in the cytochrome b6f complexAuthors : Yan, J. / Kurisu, G. / Cramer, W.A. History Deposition Sep 7, 2005 Deposition site / Processing site Revision 1.0 Dec 13, 2005 Provider / Type Revision 1.1 Apr 30, 2008 Group Revision 1.2 Jul 13, 2011 Group Revision 1.3 Feb 26, 2014 Group / OtherRevision 2.0 Oct 23, 2024 Group Data collection / Database references ... Data collection / Database references / Derived calculations / Non-polymer description / Structure summary Category chem_comp / chem_comp_atom ... chem_comp / chem_comp_atom / chem_comp_bond / database_2 / pdbx_entry_details / pdbx_modification_feature / pdbx_struct_conn_angle / pdbx_validate_chiral / struct_conn / struct_conn_type / struct_ref_seq_dif / struct_site Item _chem_comp.formula / _chem_comp.pdbx_synonyms ... _chem_comp.formula / _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_struct_conn_angle.ptnr1_auth_asym_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_asym_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr2_auth_asym_id / _pdbx_struct_conn_angle.ptnr2_auth_comp_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr2_label_atom_id / _pdbx_struct_conn_angle.ptnr2_label_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_asym_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.conn_type_id / _struct_conn.id / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_conn_type.id / _struct_ref_seq_dif.details / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id Revision 3.0 Dec 24, 2025 Group Advisory / Atomic model ... Advisory / Atomic model / Data collection / Derived calculations / Non-polymer description / Structure summary Category atom_site / chem_comp ... atom_site / chem_comp / chem_comp_atom / chem_comp_bond / database_PDB_caveat / entity / pdbx_entity_nonpoly / pdbx_modification_feature / pdbx_nonpoly_scheme / pdbx_struct_conn_angle / pdbx_validate_close_contact / struct_asym / struct_conn / struct_site / struct_site_gen Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.label_entity_id / _atom_site.type_symbol / _chem_comp.formula / _chem_comp.formula_weight / _chem_comp.id / _chem_comp.mon_nstd_flag / _chem_comp.name / _chem_comp.pdbx_synonyms / _chem_comp.type / _pdbx_modification_feature.auth_comp_id / _pdbx_modification_feature.label_comp_id / _pdbx_modification_feature.ref_comp_id / _pdbx_nonpoly_scheme.entity_id / _pdbx_nonpoly_scheme.mon_id / _pdbx_nonpoly_scheme.pdb_mon_id / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr2_auth_comp_id / _pdbx_struct_conn_angle.ptnr2_label_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_validate_close_contact.auth_comp_id_2 / _struct_asym.entity_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_label_comp_id / _struct_site.details / _struct_site.pdbx_auth_comp_id / _struct_site_gen.auth_comp_id / _struct_site_gen.label_comp_id
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Mastigocladus laminosus (bacteria)
Mastigocladus laminosus (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 3.8 Å
SYNCHROTRON / Resolution: 3.8 Å  Authors
Authors Citation
Citation Journal: Proc.Natl.Acad.Sci.USA / Year: 2006
Journal: Proc.Natl.Acad.Sci.USA / Year: 2006 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2d2c.cif.gz
2d2c.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2d2c.ent.gz
pdb2d2c.ent.gz PDB format
PDB format 2d2c.json.gz
2d2c.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/d2/2d2c
https://data.pdbj.org/pub/pdb/validation_reports/d2/2d2c ftp://data.pdbj.org/pub/pdb/validation_reports/d2/2d2c
ftp://data.pdbj.org/pub/pdb/validation_reports/d2/2d2c
 Links
Links Assembly
Assembly
 Components
Components Mastigocladus laminosus (bacteria) / References: UniProt: P83791
Mastigocladus laminosus (bacteria) / References: UniProt: P83791 Mastigocladus laminosus (bacteria) / References: UniProt: P83793
Mastigocladus laminosus (bacteria) / References: UniProt: P83793 Mastigocladus laminosus (bacteria)
Mastigocladus laminosus (bacteria) Mastigocladus laminosus (bacteria) / References: UniProt: P83792
Mastigocladus laminosus (bacteria) / References: UniProt: P83792 Mastigocladus laminosus (bacteria) / References: UniProt: P83795
Mastigocladus laminosus (bacteria) / References: UniProt: P83795 Mastigocladus laminosus (bacteria) / References: UniProt: P83796
Mastigocladus laminosus (bacteria) / References: UniProt: P83796 Mastigocladus laminosus (bacteria) / References: UniProt: P83797
Mastigocladus laminosus (bacteria) / References: UniProt: P83797 Mastigocladus laminosus (bacteria) / References: UniProt: P83798
Mastigocladus laminosus (bacteria) / References: UniProt: P83798
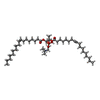
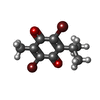










 X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 19-ID / Wavelength: 1 Å
/ Beamline: 19-ID / Wavelength: 1 Å Processing
Processing Movie
Movie Controller
Controller























 PDBj
PDBj



















