+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1yjj | ||||||
|---|---|---|---|---|---|---|---|
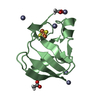
| Title | RDC-refined Solution NMR structure of oxidized putidaredoxin | ||||||
 Components Components | Putidaredoxin | ||||||
 Keywords Keywords | ELECTRON TRANSPORT / ferredoxin / [2Fe-2S] / redox / iron-sulfur / electron transfer / cytochrome P450cam | ||||||
| Function / homology |  Function and homology information Function and homology informationP450-containing electron transport chain / 2 iron, 2 sulfur cluster binding / electron transfer activity / metal ion binding / cytosol Similarity search - Function | ||||||
| Biological species |  Pseudomonas putida (bacteria) Pseudomonas putida (bacteria) | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
 Authors Authors | Jain, N.U. / Tjioe, E. / Savidor, A. / Boulie, J. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2005 Journal: Biochemistry / Year: 2005Title: Redox-dependent structural differences in putidaredoxin derived from homologous structure refinement via residual dipolar couplings. Authors: Jain, N.U. / Tjioe, E. / Savidor, A. / Boulie, J. #1:  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: A refined model for the solution structure of oxidized putidaredoxin Authors: Pochapsky, T.C. / Jain, N.U. / Kuti, M. / Lyons, T.A. / Heymont, J. #2:  Journal: Structure / Year: 1998 Journal: Structure / Year: 1998Title: New aspects of electron transfer revealed by the crystal structure of a truncated bovine adrenodoxin, ADX(4-108) Authors: MULLER, A. / MULLER, J.J. / MULLER, Y.A. / UHLMANN, H. / BERNHARDT, R. / HEINEMANN, U. #3:  Journal: Biochemistry / Year: 1994 Journal: Biochemistry / Year: 1994Title: An NMR-derived model for the solution structure of oxidized putidaredoxin, a 2Fe-2S Ferredoxin from pseudomonas Authors: POCHAPSKY, T.C. / YE, X.M. / RATNASWAMY, G. / LYONS, T.A. #4:  Journal: J.Mol.Biol. / Year: 2003 Journal: J.Mol.Biol. / Year: 2003Title: Crystal structure of putidaredoxin, the [2Fe-2S] component of the P450cam monooxygenase system from Pseudomonas putida Authors: Sevrioukova, I.F. / Garcia, C. / Li, H. / Bhaskar, B. / Poulos, T.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1yjj.cif.gz 1yjj.cif.gz | 463.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1yjj.ent.gz pdb1yjj.ent.gz | 384.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1yjj.json.gz 1yjj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/yj/1yjj https://data.pdbj.org/pub/pdb/validation_reports/yj/1yjj ftp://data.pdbj.org/pub/pdb/validation_reports/yj/1yjj ftp://data.pdbj.org/pub/pdb/validation_reports/yj/1yjj | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 11428.911 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Pseudomonas putida (bacteria) / Gene: camB / Plasmid: pkM536 / Production host: Pseudomonas putida (bacteria) / Gene: camB / Plasmid: pkM536 / Production host:  |
|---|---|
| #2: Chemical | ChemComp-FES / |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||
| NMR details | Text: The structures were refined against 15N-1H RDC restraints using previously published NOE and dihedral angle restraints for Pdx. The restraints for the metal binding loop (residues 36-48,85-86) ...Text: The structures were refined against 15N-1H RDC restraints using previously published NOE and dihedral angle restraints for Pdx. The restraints for the metal binding loop (residues 36-48,85-86) were modeled from bovine adrenodoxin crystal structure. |
- Sample preparation
Sample preparation
| Details | Contents: 1 mM oxidized putidaredoxin 15N; 50 mM Tris-Cl buffer, 50 mM KCl, pH 7.4 Solvent system: 90% H2O/10% D2O |
|---|---|
| Sample conditions | Ionic strength: 50 mM KCl / pH: 7.4 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| NMR spectrometer | Type: Varian INOVA / Manufacturer: Varian / Model: INOVA / Field strength: 600 MHz |
- Processing
Processing
| NMR software | Name:  Xplor-NIH / Version: 2.9.6 / Developer: Schwieters, Kuszewski, Tjandra, Clore / Classification: refinement Xplor-NIH / Version: 2.9.6 / Developer: Schwieters, Kuszewski, Tjandra, Clore / Classification: refinement |
|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 Details: The structures are based on 1239 NOE restriants, 160 dihedral angle restraints and 155 RDC restraints, 28 paramagnetic distance restraints from relaxation measurements and 18 distance ...Details: The structures are based on 1239 NOE restriants, 160 dihedral angle restraints and 155 RDC restraints, 28 paramagnetic distance restraints from relaxation measurements and 18 distance restriants from hydrogen bonds |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 50 / Conformers submitted total number: 15 |
 Movie
Movie Controller
Controller














 PDBj
PDBj










