+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1wp8 | ||||||
|---|---|---|---|---|---|---|---|
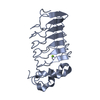
| Title | crystal structure of Hendra Virus fusion core | ||||||
 Components Components | Fusion glycoprotein F0,Fusion glycoprotein F0 | ||||||
 Keywords Keywords | VIRAL PROTEIN / Hendra Virus / Fusion Core / heptad repeat | ||||||
| Function / homology |  Function and homology information Function and homology informationhost cell surface / fusion of virus membrane with host plasma membrane / viral envelope / symbiont entry into host cell / host cell plasma membrane / virion membrane Similarity search - Function | ||||||
| Biological species |  Hendra virus Hendra virus | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 2.2 Å MAD / Resolution: 2.2 Å | ||||||
 Authors Authors | Xu, Y. / Liu, Y. / Lou, Z. / Su, N. / Bai, Z. / Gao, G.F. / Rao, Z. | ||||||
 Citation Citation |  Journal: FEBS J. / Year: 2006 Journal: FEBS J. / Year: 2006Title: Crystal structures of Nipah and Hendra virus fusion core proteins Authors: Lou, Z. / Xu, Y. / Xiang, K. / Su, N. / Qin, L. / Li, X. / Gao, G.F. / Bartlam, M. / Rao, Z. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1wp8.cif.gz 1wp8.cif.gz | 50.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1wp8.ent.gz pdb1wp8.ent.gz | 37.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1wp8.json.gz 1wp8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/wp/1wp8 https://data.pdbj.org/pub/pdb/validation_reports/wp/1wp8 ftp://data.pdbj.org/pub/pdb/validation_reports/wp/1wp8 ftp://data.pdbj.org/pub/pdb/validation_reports/wp/1wp8 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9748.781 Da / Num. of mol.: 3 / Fragment: UNP residues 137-178,UNP residues 453-485 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Hendra virus / Genus: Henipavirus / Plasmid: pET 22b / Species (production host): Escherichia coli / Production host: Hendra virus / Genus: Henipavirus / Plasmid: pET 22b / Species (production host): Escherichia coli / Production host:  #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.7 Å3/Da / Density % sol: 29.62 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: PEG4000, pH 6.5, VAPOR DIFFUSION, HANGING DROP, temperature 291K |
-Data collection
| Diffraction source | Source:  SYNCHROTRON / Site: BSRF SYNCHROTRON / Site: BSRF  / Beamline: 3W1A / Wavelength: 0.9799, 0.9801, 0.950 / Beamline: 3W1A / Wavelength: 0.9799, 0.9801, 0.950 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Date: Nov 12, 2003 | ||||||||||||
| Radiation | Protocol: MAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||
| Radiation wavelength |
| ||||||||||||
| Reflection | Resolution: 2.2→35 Å / Num. all: 9875 / Num. obs: 9875 | ||||||||||||
| Reflection shell | Resolution: 2.2→2.25 Å |
- Processing
Processing
| Software |
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 2.2→35 Å MAD / Resolution: 2.2→35 Å
| |||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→35 Å
|
 Movie
Movie Controller
Controller










 PDBj
PDBj


