[English] 日本語
 Yorodumi
Yorodumi- PDB-1uj0: Crystal Structure of STAM2 SH3 domain in complex with a UBPY-deri... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1uj0 | ||||||
|---|---|---|---|---|---|---|---|
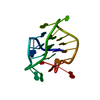
| Title | Crystal Structure of STAM2 SH3 domain in complex with a UBPY-derived peptide | ||||||
 Components Components |
| ||||||
 Keywords Keywords | SIGNALING PROTEIN/SIGNALING PROTEIN / STAM / SH3 / deubiquitinating enzyme / Grb2 / Gads / PXXP / Hrs / endocytosis / early endosome / SIGNALING PROTEIN-SIGNALING PROTEIN COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationRHOU GTPase cycle / Downregulation of ERBB2:ERBB3 signaling / Endosomal Sorting Complex Required For Transport (ESCRT) / Negative regulation of MET activity / EGFR downregulation / Regulation of FZD by ubiquitination / negative regulation of lysosomal protein catabolic process / acrosomal membrane / Cargo recognition for clathrin-mediated endocytosis / Clathrin-mediated endocytosis ...RHOU GTPase cycle / Downregulation of ERBB2:ERBB3 signaling / Endosomal Sorting Complex Required For Transport (ESCRT) / Negative regulation of MET activity / EGFR downregulation / Regulation of FZD by ubiquitination / negative regulation of lysosomal protein catabolic process / acrosomal membrane / Cargo recognition for clathrin-mediated endocytosis / Clathrin-mediated endocytosis / regulation of protein catabolic process at postsynapse, modulating synaptic transmission / multivesicular body assembly / protein K48-linked deubiquitination / endosome organization / Ub-specific processing proteases / K48-linked deubiquitinase activity / protein K63-linked deubiquitination / positive regulation of amyloid fibril formation / K63-linked deubiquitinase activity / mitotic cytokinesis / cellular response to dexamethasone stimulus / endocytic vesicle / protein deubiquitination / phosphatidylinositol binding / macroautophagy / ubiquitin binding / regulation of protein stability / cellular response to nerve growth factor stimulus / SH3 domain binding / positive regulation of canonical Wnt signaling pathway / regulation of protein localization / protein transport / early endosome membrane / midbody / Ras protein signal transduction / early endosome / cysteine-type deubiquitinase activity / ubiquitinyl hydrolase 1 / endosome membrane / postsynaptic density / glutamatergic synapse / signal transduction / proteolysis / nucleoplasm / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | ||||||
 Authors Authors | Kaneko, T. / Kumasaka, T. / Ganbe, T. / Sato, T. / Miyazawa, K. / Kitamura, N. / Tanaka, N. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2003 Journal: J.Biol.Chem. / Year: 2003Title: Structural insight into modest binding of a non-PXXP ligand to the signal transducing adaptor molecule-2 Src homology 3 domain. Authors: Kaneko, T. / Kumasaka, T. / Ganbe, T. / Sato, T. / Miyazawa, K. / Kitamura, N. / Tanaka, N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1uj0.cif.gz 1uj0.cif.gz | 28.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1uj0.ent.gz pdb1uj0.ent.gz | 18.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1uj0.json.gz 1uj0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/uj/1uj0 https://data.pdbj.org/pub/pdb/validation_reports/uj/1uj0 ftp://data.pdbj.org/pub/pdb/validation_reports/uj/1uj0 ftp://data.pdbj.org/pub/pdb/validation_reports/uj/1uj0 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 7059.673 Da / Num. of mol.: 1 / Fragment: RESIDUES 200-261 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #2: Protein/peptide | Mass: 1284.483 Da / Num. of mol.: 1 / Fragment: RESIDUES 699-709 / Source method: obtained synthetically Details: The peptide was chemically synthesized. The sequence of the peptide is naturally found in Mus musculus (mouse). References: GenBank: 11414862, UniProt: Q80U87*PLUS |
| #3: Chemical | ChemComp-PO4 / |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.29 Å3/Da / Density % sol: 45.97 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop Details: 0.08M Sodium Phosphate monobasic, 1.92M Potassium phosphate dibasic, VAPOR DIFFUSION, HANGING DROP, temperature 277K | ||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 25 ℃ / Method: vapor diffusion | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SPring-8 SPring-8  / Beamline: BL26B1 / Wavelength: 1 Å / Beamline: BL26B1 / Wavelength: 1 Å |
| Detector | Type: RIGAKU / Detector: IMAGE PLATE / Date: Oct 2, 2002 |
| Radiation | Monochromator: SI / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→34.1 Å / Num. obs: 8501 / % possible obs: 97.9 % / Observed criterion σ(I): 6 |
| Reflection shell | Resolution: 1.7→1.79 Å / % possible all: 96.8 |
| Reflection | *PLUS Rmerge(I) obs: 0.064 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.7→34.11 Å / Cor.coef. Fo:Fc: 0.933 / Cor.coef. Fo:Fc free: 0.932 / SU B: 2.525 / SU ML: 0.083 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.133 / ESU R Free: 0.116 / Stereochemistry target values: MAXIMUM LIKELIHOOD MOLECULAR REPLACEMENT / Resolution: 1.7→34.11 Å / Cor.coef. Fo:Fc: 0.933 / Cor.coef. Fo:Fc free: 0.932 / SU B: 2.525 / SU ML: 0.083 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.133 / ESU R Free: 0.116 / Stereochemistry target values: MAXIMUM LIKELIHOOD
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 25.802 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→34.11 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.698→1.742 Å / Total num. of bins used: 20 /
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS % reflection Rfree: 10 % / Rfactor Rfree: 0.23 / Rfactor Rwork: 0.215 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller








 PDBj
PDBj








