[English] 日本語
 Yorodumi
Yorodumi- PDB-1tvq: Crystal Structure of Apo Chicken Liver Basic Fatty Acid Binding P... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tvq | ||||||
|---|---|---|---|---|---|---|---|
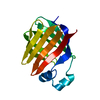
| Title | Crystal Structure of Apo Chicken Liver Basic Fatty Acid Binding Protein (or Bile Acid Binding Protein) | ||||||
 Components Components | Fatty acid-binding protein | ||||||
 Keywords Keywords | LIPID BINDING PROTEIN / Lb-FABP / FABP / BABP / INTRACELLULAR BILE ACID-BINDING PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIR / Resolution: 2 Å MIR / Resolution: 2 Å | ||||||
 Authors Authors | Nichesola, D. / Perduca, M. / Capaldi, S. / Carrizo, M.E. / Righetti, P.G. / Monaco, H.L. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2004 Journal: Biochemistry / Year: 2004Title: Crystal structure of chicken liver basic Fatty Acid-binding protein complexed with cholic acid Authors: Nichesola, D. / Perduca, M. / Capaldi, S. / Carrizo, M.E. / Righetti, P.G. / Monaco, H.L. #1: Journal: MOL.CELL.BIOCHEM. / Year: 1990 Title: Crystal Structure of Chicken Liver Basic Fatty Acid-Binding Protein at 2.7 A Resolution Authors: Scapin, G. / Spadon, P. / Mammi, M. / Zanotti, G. / Monaco, H.L. #2: Journal: FEBS Lett. / Year: 1988 Title: Chicken liver basic fatty acid-binding protein (pI = 9.0). Purification, crystallization and preliminary X-ray data Authors: Scapin, G. / Spadon, P. / Pengo, L. / Mammi, M. / Zanotti, G. / Monaco, H.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tvq.cif.gz 1tvq.cif.gz | 38.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tvq.ent.gz pdb1tvq.ent.gz | 26.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tvq.json.gz 1tvq.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tv/1tvq https://data.pdbj.org/pub/pdb/validation_reports/tv/1tvq ftp://data.pdbj.org/pub/pdb/validation_reports/tv/1tvq ftp://data.pdbj.org/pub/pdb/validation_reports/tv/1tvq | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14100.177 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  |
|---|---|
| #2: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.8 Å3/Da / Density % sol: 55.8 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: microgravity / pH: 7.5 Details: IMIDAZOLE, PEG 6000, pH 7.5, MICROGRAVITY, temperature 298.0K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ELETTRA ELETTRA  / Beamline: 5.2R / Wavelength: 1 Å / Beamline: 5.2R / Wavelength: 1 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Sep 12, 2001 / Details: Three-segment Pt-coated toroidal |
| Radiation | Monochromator: Double Crystal (Si111, Si220) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2→30 Å / Num. all: 60486 / Num. obs: 60486 / % possible obs: 99.3 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 5.5 % / Biso Wilson estimate: 22.84 Å2 / Rsym value: 0.047 |
| Reflection shell | Resolution: 2→2.11 Å / Rsym value: 0.047 / % possible all: 100 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIR / Resolution: 2→29.54 Å / Rfactor Rfree error: 0.008 / Data cutoff high absF: 8119382.96 / Data cutoff low absF: 0 / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber MIR / Resolution: 2→29.54 Å / Rfactor Rfree error: 0.008 / Data cutoff high absF: 8119382.96 / Data cutoff low absF: 0 / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24.81 Å2 | |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→29.54 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.07 Å /
|
 Movie
Movie Controller
Controller












 PDBj
PDBj


