+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qtx | ||||||
|---|---|---|---|---|---|---|---|
| Title | THE 1.65 ANGSTROM STRUCTURE OF CALMODULIN RS20 PEPTIDE COMPLEX | ||||||
 Components Components |
| ||||||
 Keywords Keywords | SIGNALING PROTEIN / CALMODULIN / CALCIUM BINDING / HELIX-LOOP-HELIX / SIGNALING / COMPLEX (CALCIUM- BINDING PROTEIN-PEPTIDE) | ||||||
| Function / homology |  Function and homology information Function and homology informationtonic smooth muscle contraction / myosin-light-chain kinase / myosin light chain kinase activity / positive regulation of sarcomere organization / positive regulation of striated muscle contraction / myosin II complex / cardiac muscle cell development / cleavage furrow / heart morphogenesis / stress fiber ...tonic smooth muscle contraction / myosin-light-chain kinase / myosin light chain kinase activity / positive regulation of sarcomere organization / positive regulation of striated muscle contraction / myosin II complex / cardiac muscle cell development / cleavage furrow / heart morphogenesis / stress fiber / lamellipodium / calmodulin binding / calcium ion binding / ATP binding / metal ion binding / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 1.65 Å MOLECULAR REPLACEMENT / Resolution: 1.65 Å | ||||||
 Authors Authors | Weigand, S. / Anderson, W.F. | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: High Resolution Structure of a Calmodulin Rs20 Peptide Complex Authors: Weigand, S. / Shuvalova, L. / Lukas, T.J. / Mirzoeva, S. / Watterson, D.M. / Anderson, W.F. #1:  Journal: Biochemistry / Year: 1999 Journal: Biochemistry / Year: 1999Title: Analysis of the Functional Coupling between Calmodulin S Calcium Binding and Peptide Recognition Properties Authors: Mirzoeva, S. / Weigand, S. / Lukas, T.J. / Shuvalova, L. / Anderson, W.F. / Watterson, D.M. #2:  Journal: Science / Year: 1992 Journal: Science / Year: 1992Title: Target Enzyme Recognition by Calmodulin: 2.4 A Structure of a Calmodulin- Peptide Complex Authors: Meador, W.E. / Means, A.R. / Quiocho, F.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qtx.cif.gz 1qtx.cif.gz | 59.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qtx.ent.gz pdb1qtx.ent.gz | 43 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qtx.json.gz 1qtx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qt/1qtx https://data.pdbj.org/pub/pdb/validation_reports/qt/1qtx ftp://data.pdbj.org/pub/pdb/validation_reports/qt/1qtx ftp://data.pdbj.org/pub/pdb/validation_reports/qt/1qtx | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 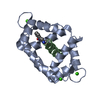 1qs7C  1vrmS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 16642.271 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   | ||||
|---|---|---|---|---|---|
| #2: Protein/peptide | Mass: 2299.686 Da / Num. of mol.: 1 Fragment: CALMODULIN BINDING REGION FROM SMOOTH MUSCLE/NONMUSCLE MYOSIN LIGHT CHAIN KINASE Source method: obtained synthetically Details: PEPTIDE ANALOG OF THE CALMODULIN RECOGNITION REGION OF CHICKEN SMOOTH MUSCLE/ NONMUSCLE MYOSIN LIGHT CHAIN KINASE References: UniProt: P11799 | ||||
| #3: Chemical | ChemComp-CA / #4: Water | ChemComp-HOH / | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 3 X-RAY DIFFRACTION / Number of used crystals: 3 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.11 Å3/Da / Density % sol: 41.8 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 4.6 Details: 20% (W/V) PEG 8000, 100 MM SODIUM ACETATE 5 MM CALCIUM CHLORIDE 0.01% (W/V) SODIUM AZIDE PH = 4.6, VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 | ||||||||||||||||||||
| Detector |
| ||||||||||||||||||||
| Radiation | Monochromator: FILTER / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 | ||||||||||||||||||||
| Reflection | Resolution: 1.53→30 Å / Num. all: 166836 / Num. obs: 19966 / % possible obs: 79.3 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -3 / Redundancy: 8.4 % / Biso Wilson estimate: 19.6 Å2 / Rmerge(I) obs: 0.105 / Net I/σ(I): 19.6 | ||||||||||||||||||||
| Reflection shell | Resolution: 1.65→1.68 Å / Redundancy: 5.5 % / Rmerge(I) obs: 0.221 / Mean I/σ(I) obs: 6.9 / % possible all: 59.4 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1VRM CHAIN A AND CHAIN B Resolution: 1.65→29.595 Å / Rfactor Rfree error: 0.005 / Data cutoff high absF: 1176942.78 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): -3 Stereochemistry target values: ENGH, R.A. AND HUBER, R. (1991). ACTA CRYST. A47, 392-400.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 22.2 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.65→29.595 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.65→1.69 Å / Rfactor Rfree error: 0.028 / Total num. of bins used: 16
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller














 PDBj
PDBj







