+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1pjg | ||||||
|---|---|---|---|---|---|---|---|
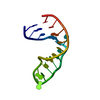
| Title | RNA/DNA Hybrid Decamer of CAAAGAAAAG/CTTTTCTTTG | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA-RNA HYBRID / RNA/DNA hybrid / Polypurine Tract of HIV-1 / Moleuclar Replacement / sugar conformation / DNA-RNA COMPLEX | ||||||
| Function / homology | DNA / RNA Function and homology information Function and homology information | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.15 Å MOLECULAR REPLACEMENT / Resolution: 1.15 Å | ||||||
 Authors Authors | Kopka, M.L. / Lavelle, L. / Han, G.W. / Ng, H.-L. / Dickerson, R.E. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2003 Journal: J.Mol.Biol. / Year: 2003Title: An unusual sugar conformation in the structure of an RNA/DNA decamer of the polypurine tract may affect recognition by RNase H. Authors: Kopka, M.L. / Lavelle, L. / Han, G.W. / Ng, H.L. / Dickerson, R.E. #1:  Journal: Acta Crystallogr.,Sect.D / Year: 2001 Journal: Acta Crystallogr.,Sect.D / Year: 2001Title: Direct Methods Determination of an RNA/DNA Hybrid Decamer at 1.15A Resolution Authors: Han, G.W. #2:  Journal: To be Published / Year: 2003 Journal: To be Published / Year: 2003Title: Crystal Structure of an RNA/DNA Hybrid Reveals Novel Intermolecular Intercalation: Dimer Formation by Base-pair Swapping Authors: Han, G.W. / Kopka, M.L. / Langs, D. / Sawaya, M.R. / Dickerson, R.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1pjg.cif.gz 1pjg.cif.gz | 46.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1pjg.ent.gz pdb1pjg.ent.gz | 33.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1pjg.json.gz 1pjg.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pj/1pjg https://data.pdbj.org/pub/pdb/validation_reports/pj/1pjg ftp://data.pdbj.org/pub/pdb/validation_reports/pj/1pjg ftp://data.pdbj.org/pub/pdb/validation_reports/pj/1pjg | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1pjoC C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 3255.076 Da / Num. of mol.: 1 / Fragment: Polypurine Tract of HIV-1 / Source method: obtained synthetically | ||
|---|---|---|---|
| #2: DNA chain | Mass: 2991.961 Da / Num. of mol.: 1 / Source method: obtained synthetically | ||
| #3: Chemical | ChemComp-CA / #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.93 Å3/Da / Density % sol: 36.38 % | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 278 K / Method: vapor diffusion, hanging drop / pH: 6.8 Details: calcium acetate, cacodylate, beta-octylglucoside, spermidine hydrochloride, 2-methyl,2,4-pentanediol, pH 6.8, VAPOR DIFFUSION, HANGING DROP, temperature 278K | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion | ||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 123 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X8C / Wavelength: 1.072 Å / Beamline: X8C / Wavelength: 1.072 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Oct 27, 1999 / Details: mirrors |
| Radiation | Monochromator: mirrors / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.072 Å / Relative weight: 1 |
| Reflection | Resolution: 1.15→50 Å / Num. obs: 17368 / % possible obs: 96.2 % / Observed criterion σ(I): 4.7 / Biso Wilson estimate: 8.17 Å2 / Rmerge(I) obs: 0.081 / Net I/σ(I): 29.6 |
| Reflection shell | Resolution: 1.15→1.18 Å / Rmerge(I) obs: 0.261 / Mean I/σ(I) obs: 11.09 / Num. unique all: 882 / % possible all: 93.9 |
| Reflection | *PLUS Lowest resolution: 50 Å / % possible obs: 96 % |
| Reflection shell | *PLUS % possible obs: 93.9 % |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: NDB ID AH0001 Resolution: 1.15→8 Å / σ(F): 0
| |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 10.71 Å2 | |||||||||||||||||||||||||
| Refine analyze | Occupancy sum hydrogen: 1 / Occupancy sum non hydrogen: 1 | |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.15→8 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.15→1.18 Å /
| |||||||||||||||||||||||||
| Software | *PLUS Name: SHELXL / Version: 97 / Classification: refinement | |||||||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.158 / Rfactor Rwork: 0.127 | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller







 PDBj
PDBj


