[English] 日本語
 Yorodumi
Yorodumi- PDB-1ow8: Paxillin LD2 motif bound to the Focal Adhesion Targeting (FAT) do... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ow8 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Paxillin LD2 motif bound to the Focal Adhesion Targeting (FAT) domain of the Focal Adhesion Kinase | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSFERASE / 4 helical bundle / amphiphatic helix | ||||||
| Function / homology |  Function and homology information Function and homology informationnetrin-activated signaling pathway / regulation of substrate adhesion-dependent cell spreading / Regulation of cytoskeletal remodeling and cell spreading by IPP complex components / Localization of the PINCH-ILK-PARVIN complex to focal adhesions / regulation of endothelial cell migration / regulation of epithelial cell migration / Regulation of MITF-M-dependent genes involved in extracellular matrix, focal adhesion and epithelial-to-mesenchymal transition / detection of muscle stretch / neuropilin binding / positive regulation of cell-cell adhesion mediated by integrin ...netrin-activated signaling pathway / regulation of substrate adhesion-dependent cell spreading / Regulation of cytoskeletal remodeling and cell spreading by IPP complex components / Localization of the PINCH-ILK-PARVIN complex to focal adhesions / regulation of endothelial cell migration / regulation of epithelial cell migration / Regulation of MITF-M-dependent genes involved in extracellular matrix, focal adhesion and epithelial-to-mesenchymal transition / detection of muscle stretch / neuropilin binding / positive regulation of cell-cell adhesion mediated by integrin / vinculin binding / JUN kinase binding / signal complex assembly / negative regulation of cell adhesion mediated by integrin / positive regulation of macrophage proliferation / DCC mediated attractive signaling / Signal regulatory protein family interactions / positive regulation of fibroblast migration / microtubule associated complex / MET activates PTK2 signaling / growth hormone receptor signaling pathway / positive regulation of ubiquitin-dependent protein catabolic process / regulation of focal adhesion assembly / negative regulation of cell-cell adhesion / Apoptotic cleavage of cellular proteins / p130Cas linkage to MAPK signaling for integrins / positive regulation of wound healing / regulation of osteoblast differentiation / Fc-gamma receptor signaling pathway involved in phagocytosis / regulation of cytoskeleton organization / positive regulation of macrophage chemotaxis / establishment of cell polarity / vascular endothelial cell response to oscillatory fluid shear stress / GRB2:SOS provides linkage to MAPK signaling for Integrins / negative regulation of anoikis / positive regulation of epithelial cell migration / Estrogen-dependent nuclear events downstream of ESR-membrane signaling / vascular endothelial growth factor receptor signaling pathway / ephrin receptor signaling pathway / Smooth Muscle Contraction / GAB1 signalosome / RHO GTPases Activate WASPs and WAVEs / positive regulation of epithelial to mesenchymal transition / endothelial cell migration / regulation of cell adhesion / heart morphogenesis / PTK6 Regulates RHO GTPases, RAS GTPase and MAP kinases / positive regulation of stress fiber assembly / Integrin signaling / stress fiber / EPHB-mediated forward signaling / substrate adhesion-dependent cell spreading / protein tyrosine phosphatase activity / transforming growth factor beta receptor signaling pathway / NCAM signaling for neurite out-growth / SH2 domain binding / ciliary tip / Turbulent (oscillatory, disturbed) flow shear stress activates signaling by PIEZO1 and integrins in endothelial cells / molecular function activator activity / placenta development / axon guidance / integrin-mediated signaling pathway / FCGR3A-mediated phagocytosis / non-specific protein-tyrosine kinase / cellular response to reactive oxygen species / cell motility / non-membrane spanning protein tyrosine kinase activity / Regulation of actin dynamics for phagocytic cup formation / beta-catenin binding / VEGFA-VEGFR2 Pathway / integrin binding / epidermal growth factor receptor signaling pathway / cell-cell junction / regulation of cell shape / regulation of cell population proliferation / cell migration / lamellipodium / RAF/MAP kinase cascade / actin binding / protein tyrosine kinase activity / angiogenesis / cell cortex / protein phosphatase binding / cytoskeleton / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / cell adhesion / Extra-nuclear estrogen signaling / cilium / positive regulation of cell migration / ciliary basal body / focal adhesion / positive regulation of cell population proliferation / centrosome / protein kinase binding / negative regulation of apoptotic process / perinuclear region of cytoplasm / signal transduction / ATP binding / metal ion binding / nucleus Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.85 Å MOLECULAR REPLACEMENT / Resolution: 2.85 Å | ||||||
 Authors Authors | Hoellerer, M.K. / Noble, M.E.M. / Labesse, G. / Werner, J.M. / Arold, S.T. | ||||||
 Citation Citation |  Journal: Structure / Year: 2003 Journal: Structure / Year: 2003Title: Molecular Recognition of Paxillin LD Motifs by the Focal Adhesion Targeting Domain Authors: Hoellerer, M.K. / Noble, M.E.M. / Labesse, G. / Werner, J.M. / Arold, S.T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ow8.cif.gz 1ow8.cif.gz | 95.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ow8.ent.gz pdb1ow8.ent.gz | 75.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ow8.json.gz 1ow8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ow/1ow8 https://data.pdbj.org/pub/pdb/validation_reports/ow/1ow8 ftp://data.pdbj.org/pub/pdb/validation_reports/ow/1ow8 ftp://data.pdbj.org/pub/pdb/validation_reports/ow/1ow8 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 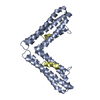 1ow6C  1ow7C  1k05S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 17836.539 Da / Num. of mol.: 3 / Fragment: Focal Adhesion Targeting Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PTK2 OR FAK1 OR FAK / Plasmid: pGEX-6P2 / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Gene: PTK2 OR FAK1 OR FAK / Plasmid: pGEX-6P2 / Species (production host): Escherichia coli / Production host:  #2: Protein/peptide | Mass: 1542.755 Da / Num. of mol.: 2 / Fragment: Paxillin LD2 motif / Source method: obtained synthetically / Details: Sequence derived from human paxillin / References: UniProt: P49023*PLUS #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 4.86 Å3/Da / Density % sol: 74.51 % | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 292 K / Method: vapor diffusion, hanging drop / pH: 6.3 Details: sodium chloride, glycerol, Hepes, pH 6.3, VAPOR DIFFUSION, HANGING DROP, temperature 292K | ||||||||||||||||||||||||||||
| Crystal grow | *PLUS Temperature: 18 ℃ / Method: vapor diffusion, hanging drop | ||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: BM30A / Wavelength: 0.98 Å / Beamline: BM30A / Wavelength: 0.98 Å |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Jun 30, 2002 Details: two crystals monochromator between two cylindrical parabolic mirrors |
| Radiation | Monochromator: silicon / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.98 Å / Relative weight: 1 |
| Reflection | Resolution: 2.85→34.5 Å / Num. all: 22456 / Num. obs: 22456 / % possible obs: 99.6 % / Redundancy: 3.7 % / Biso Wilson estimate: 82.6 Å2 / Rmerge(I) obs: 0.12 / Rsym value: 0.1 / Net I/σ(I): 5.1 |
| Reflection shell | Resolution: 2.85→2.95 Å / Redundancy: 3.2 % / Rmerge(I) obs: 0.81 / Mean I/σ(I) obs: 1.2 / Num. unique all: 2159 / Rsym value: 0.68 / % possible all: 99.3 |
| Reflection | *PLUS Num. measured all: 82912 / Rmerge(I) obs: 0.104 |
| Reflection shell | *PLUS % possible obs: 99.3 % / Rmerge(I) obs: 0.68 / Mean I/σ(I) obs: 1.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDBentry 1K05 Resolution: 2.85→34.5 Å / Cor.coef. Fo:Fc: 0.94 / Cor.coef. Fo:Fc free: 0.924 / Cross valid method: THROUGHOUT / σ(F): 0 / σ(I): 0 / ESU R: 0.471 / ESU R Free: 0.324 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: The second LD2 motif (chain F, bound to molecule C) was fitted into the unbiased 3Fo-2Fc maps (CNS). Its position and orientation were inferred from homology with the well-defined other LD2 - ...Details: The second LD2 motif (chain F, bound to molecule C) was fitted into the unbiased 3Fo-2Fc maps (CNS). Its position and orientation were inferred from homology with the well-defined other LD2 - FAT interaction (chains A and C), and are supported by NMR data (see publication).
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: BABINET MODEL WITH MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 67.913 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.85→34.5 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.85→2.924 Å / Total num. of bins used: 20 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS % reflection Rfree: 7 % / Rfactor Rfree: 0.275 / Rfactor Rwork: 0.235 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller















 PDBj
PDBj















