[English] 日本語
 Yorodumi
Yorodumi- PDB-1oqx: G-2 glycovariant of human IgG Fc bound to minimized version of Pr... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1oqx | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
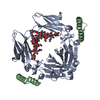
| Title | G-2 glycovariant of human IgG Fc bound to minimized version of Protein A called Z34C | ||||||||||||
 Components Components |
| ||||||||||||
 Keywords Keywords | IMMUNE SYSTEM / Antibody / Glycosylation / Oligosaccharides / IgG / Peptide complex / Z34C / Protein A | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationcomplement-dependent cytotoxicity / antibody-dependent cellular cytotoxicity / Fc-gamma receptor I complex binding / immunoglobulin complex, circulating / Classical antibody-mediated complement activation / immunoglobulin receptor binding / Initial triggering of complement / IgG immunoglobulin complex / FCGR activation / complement activation, classical pathway ...complement-dependent cytotoxicity / antibody-dependent cellular cytotoxicity / Fc-gamma receptor I complex binding / immunoglobulin complex, circulating / Classical antibody-mediated complement activation / immunoglobulin receptor binding / Initial triggering of complement / IgG immunoglobulin complex / FCGR activation / complement activation, classical pathway / Role of phospholipids in phagocytosis / antigen binding / FCGR3A-mediated IL10 synthesis / Regulation of Complement cascade / B cell receptor signaling pathway / FCGR3A-mediated phagocytosis / Regulation of actin dynamics for phagocytic cup formation / antibacterial humoral response / Interleukin-4 and Interleukin-13 signaling / blood microparticle / adaptive immune response / : / extracellular exosome / extracellular region / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human)synthetic construct (others) | ||||||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.6 Å MOLECULAR REPLACEMENT / Resolution: 2.6 Å | ||||||||||||
 Authors Authors | Raju, T.S. / Mulkerrin, M.G. / Parker, M. / De Vos, A.M. / Gazzano-Santoro, H. / Totpal, K. / Ultsch, M.H. | ||||||||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Impact of Fc Glycans on The Effector Functions Vary with the Antibody mechanism of Action Authors: Raju, T.S. / Mulkerrin, M.G. / Parker, M. / De Vos, A.M. / Gazzano-Santoro, H. / Totpal, K. / Ultsch, M.H. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1oqx.cif.gz 1oqx.cif.gz | 117.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1oqx.ent.gz pdb1oqx.ent.gz | 89.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1oqx.json.gz 1oqx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/oq/1oqx https://data.pdbj.org/pub/pdb/validation_reports/oq/1oqx ftp://data.pdbj.org/pub/pdb/validation_reports/oq/1oqx ftp://data.pdbj.org/pub/pdb/validation_reports/oq/1oqx | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1oqoC  1l6xS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Antibody | Mass: 23975.072 Da / Num. of mol.: 2 / Fragment: Fc Fragment Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  #2: Protein/peptide | Mass: 4190.682 Da / Num. of mol.: 2 / Fragment: Minimized B-domain / Source method: obtained synthetically Details: Phage Optimized Sequence from the B-Domain of Protein A Staphylococcus Aureus. Peptide Prepared using FMOC Chemistry and Wang Resin. Source: (synth.) synthetic construct (others) #3: Polysaccharide | alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2- ...alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | Source method: isolated from a genetically manipulated source #4: Polysaccharide | alpha-D-mannopyranose-(1-3)-[beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2- ...alpha-D-mannopyranose-(1-3)-[beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | Source method: isolated from a genetically manipulated source #5: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.53 Å3/Da / Density % sol: 50.05 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop / pH: 5.5 Details: PEG 550 MME, 0.1M NaOAc, 0.25M NaCl, pH 5.5, VAPOR DIFFUSION, SITTING DROP |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL9-1 / Wavelength: 1.08 Å / Beamline: BL9-1 / Wavelength: 1.08 Å |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Apr 27, 1998 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.08 Å / Relative weight: 1 |
| Reflection | Resolution: 2.6→99 Å / Num. all: 18601 / Num. obs: 18601 / % possible obs: 97.4 % / Observed criterion σ(F): -1 / Observed criterion σ(I): -1 / Redundancy: 4 % / Biso Wilson estimate: 59.3 Å2 |
| Reflection shell | Resolution: 2.59→2.68 Å / % possible all: 91.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ID 1L6X Resolution: 2.6→20 Å / Rfactor Rfree error: 0.007 / Data cutoff high absF: 10000000 / Data cutoff low absF: 0.01 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0.2 / Stereochemistry target values: Engh & Huber / Details: BULK SOLVENT MODEL USED
| ||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 55.8 Å2
| ||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.6→20 Å
| ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.6→2.76 Å / Rfactor Rfree error: 0.026 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller









 PDBj
PDBj










