[English] 日本語
 Yorodumi
Yorodumi- PDB-1mtr: HIV-1 PROTEASE COMPLEXED WITH A CYCLIC PHE-ILE-VAL PEPTIDOMIMETIC... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1mtr | ||||||
|---|---|---|---|---|---|---|---|
| Title | HIV-1 PROTEASE COMPLEXED WITH A CYCLIC PHE-ILE-VAL PEPTIDOMIMETIC INHIBITOR | ||||||
 Components Components | HIV-1 PROTEASE | ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE INHIBITOR / ASPARTYL PROTEINASE / AIDS / ASPARTYL PROTEASE / HYDROLASE-HYDROLASE INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology informationHIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / host multivesicular body / DNA integration / viral genome integration into host DNA / RNA-directed DNA polymerase / establishment of integrated proviral latency ...HIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / host multivesicular body / DNA integration / viral genome integration into host DNA / RNA-directed DNA polymerase / establishment of integrated proviral latency / viral penetration into host nucleus / RNA stem-loop binding / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / host cell / viral nucleocapsid / DNA recombination / DNA-directed DNA polymerase / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / symbiont-mediated suppression of host gene expression / viral translational frameshifting / lipid binding / symbiont entry into host cell / host cell nucleus / host cell plasma membrane / virion membrane / structural molecule activity / proteolysis / DNA binding / zinc ion binding / membrane Similarity search - Function | ||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | ||||||
| Method |  X-RAY DIFFRACTION / DIFFERENCE FOURIER / Resolution: 1.75 Å X-RAY DIFFRACTION / DIFFERENCE FOURIER / Resolution: 1.75 Å | ||||||
 Authors Authors | Wickramasinghe, W. / Begun, J. / Martin, J.L. | ||||||
 Citation Citation | Journal: J.Am.Chem.Soc. / Year: 1996 Title: Substrate-based cyclic peptidomimetics of Phe-Ile-Val that inhibit HIV-1 protease using a novel enzyme-binding mode. Authors: March, D.R. / Abbenante, G. / Bergman, D.A. / Brinkworth, R.I. / Wickramasinghe, W. / Begun, J. / Martin, J.L. / Fairlie, D.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1mtr.cif.gz 1mtr.cif.gz | 53.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1mtr.ent.gz pdb1mtr.ent.gz | 40.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1mtr.json.gz 1mtr.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  1mtr_validation.pdf.gz 1mtr_validation.pdf.gz | 494.8 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  1mtr_full_validation.pdf.gz 1mtr_full_validation.pdf.gz | 497.2 KB | Display | |
| Data in XML |  1mtr_validation.xml.gz 1mtr_validation.xml.gz | 6.4 KB | Display | |
| Data in CIF |  1mtr_validation.cif.gz 1mtr_validation.cif.gz | 9.9 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mt/1mtr https://data.pdbj.org/pub/pdb/validation_reports/mt/1mtr ftp://data.pdbj.org/pub/pdb/validation_reports/mt/1mtr ftp://data.pdbj.org/pub/pdb/validation_reports/mt/1mtr | HTTPS FTP |
-Related structure data
| Related structure data |  1cpiS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 10765.687 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Human immunodeficiency virus 1 / Genus: Lentivirus / Production host: Human immunodeficiency virus 1 / Genus: Lentivirus / Production host:  #2: Chemical | #3: Chemical | ChemComp-PI6 / [ | 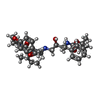  Type: peptide-like, Peptide-like / Class: Inhibitor / Mass: 596.757 Da / Num. of mol.: 1 / Source method: obtained synthetically / Formula: C33H48N4O6 Type: peptide-like, Peptide-like / Class: Inhibitor / Mass: 596.757 Da / Num. of mol.: 1 / Source method: obtained synthetically / Formula: C33H48N4O6References: tert-butyl [(1S,2S)-1-benzyl-2-hydroxy-3-{[(8S,11R)-8-[(1R)-1-methylpropyl]-7,10-dioxo-2-oxa-6,9-diazabicyclo[11.2.2]heptadeca-1(15),13,16-trien-11-yl]amino}propyl]carbamate #4: Water | ChemComp-HOH / | Sequence details | GLN 7 WAS REPLACED BY LYS TO PREVENT AUTOCLEAVAGE. LEU 33 WAS REPLACED BY ILE TO PREVENT ...GLN 7 WAS REPLACED BY LYS TO PREVENT AUTOCLEAVA | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.11 Å3/Da / Density % sol: 44 % | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 5.5 / Details: pH 5.5 | ||||||||||||||||||
| Crystal | *PLUS | ||||||||||||||||||
| Crystal grow | *PLUS Temperature: 20 ℃ / pH: 5.4 / Method: vapor diffusion | ||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 289 K |
|---|---|
| Diffraction source | Wavelength: 1.5418 |
| Detector | Type: RIGAKU RAXIS IIC / Detector: IMAGE PLATE / Date: Mar 28, 1995 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.75→50 Å / Num. obs: 16528 / % possible obs: 85.9 % / Observed criterion σ(I): 1 / Redundancy: 5.3 % / Rmerge(I) obs: 0.068 / Net I/σ(I): 10.5 |
| Reflection shell | Resolution: 1.75→2 Å / Redundancy: 2.5 % / Rmerge(I) obs: 0.219 / Mean I/σ(I) obs: 2.42 / % possible all: 69.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: DIFFERENCE FOURIER Starting model: 1CPI Resolution: 1.75→8 Å / σ(F): 2 Details: RESIDUES 50 AND 51 IN THE FLAP REGION OF BOTH SUBUNITS ARE DISORDERED AND WERE MODELLED IN TWO CONFORMATIONS. HOWEVER, THESE REGIONS OF THE STRUCTURE HAVE RELATIVELY POOR GEOMETRY.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 18.7 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.75→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Version: 3.1 / Classification: refinement X-PLOR / Version: 3.1 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller
















 PDBj
PDBj




