[English] 日本語
 Yorodumi
Yorodumi- PDB-1kl8: NMR STRUCTURAL ANALYSIS OF THE COMPLEX FORMED BETWEEN ALPHA-BUNGA... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1kl8 | ||||||
|---|---|---|---|---|---|---|---|
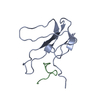
| Title | NMR STRUCTURAL ANALYSIS OF THE COMPLEX FORMED BETWEEN ALPHA-BUNGAROTOXIN AND THE PRINCIPAL ALPHA-NEUROTOXIN BINDING SEQUENCE ON THE ALPHA7 SUBUNIT OF A NEURONAL NICOTINIC ACETYLCHOLINE RECEPTOR | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TOXIN / ALPHA-BUNGAROTOXIN / NICOTINIC ACETYLCHOLINE RECEPTOR ALPHA 7 SUBUNIT / ALPHA-NEUROTOXIN / LIGAND-GATED ION CHANNELS / NMR PROTEIN-PROTEIN INTERACTIONS / PROTEIN-PEPTIDE COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationacetylcholine receptor activity / acetylcholine-gated channel complex / acetylcholine receptor inhibitor activity / chloride channel regulator activity / acetylcholine-gated monoatomic cation-selective channel activity / ion channel regulator activity / acetylcholine binding / synaptic transmission, cholinergic / negative regulation of tumor necrosis factor production / toxic substance binding ...acetylcholine receptor activity / acetylcholine-gated channel complex / acetylcholine receptor inhibitor activity / chloride channel regulator activity / acetylcholine-gated monoatomic cation-selective channel activity / ion channel regulator activity / acetylcholine binding / synaptic transmission, cholinergic / negative regulation of tumor necrosis factor production / toxic substance binding / negative regulation of cytokine production involved in inflammatory response / response to nicotine / regulation of membrane potential / cognition / positive regulation of angiogenesis / intracellular calcium ion homeostasis / calcium ion transport / amyloid-beta binding / toxin activity / monoatomic ion transmembrane transport / chemical synaptic transmission / perikaryon / response to hypoxia / positive regulation of MAPK cascade / postsynaptic membrane / neuron projection / axon / positive regulation of cell population proliferation / synapse / dendrite / signal transduction / protein homodimerization activity / extracellular region / metal ion binding / plasma membrane / cytoplasm Similarity search - Function | ||||||
| Biological species |   Bungarus multicinctus (many-banded krait) Bungarus multicinctus (many-banded krait) | ||||||
| Method | SOLUTION NMR / DISTANCE GEOMETRY DYNAMIC SIMULATED ANNEALING | ||||||
| Model type details | minimized average | ||||||
 Authors Authors | Moise, L. / Piserchio, A. / Basus, V.J. / Hawrot, E. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2002 Journal: J.Biol.Chem. / Year: 2002Title: NMR structural analysis of alpha-bungarotoxin and its complex with the principal alpha-neurotoxin-binding sequence on the alpha 7 subunit of a neuronal nicotinic acetylcholine receptor. Authors: Moise, L. / Piserchio, A. / Basus, V.J. / Hawrot, E. #1:  Journal: J.Biol.Chem. / Year: 1999 Journal: J.Biol.Chem. / Year: 1999Title: Chimeric Analysis of a Neuronal Nicotinic Acetylcholine Receptor Reveals Amino Acids Conferring Sensitivity to Alpha-Bungarotoxin. Authors: Levandoski, M.M. / Lin, Y. / Moise, L. / Mclaughlin, J.T. / Cooper, E. / Hawrot, E. #2:  Journal: J.Biol.Chem. / Year: 1991 Journal: J.Biol.Chem. / Year: 1991Title: Identification of Sequence Segments Forming the Alpha-Bungarotoxin Binding Sites on Two Nicotinic Acetylcholine Receptor Alpha Subunits from the Avian Brain. Authors: Mclane, K.E. / Wu, X.D. / Schoepfer, R. / Lindstrom, J.M. / Conti-Tronconi, B.M. #3:  Journal: Biochemistry / Year: 1988 Journal: Biochemistry / Year: 1988Title: Structural Studies of Alpha-Bungarotoxin. 1. Sequence-Specific 1H NMR Resonance Assignments. Authors: Basus, V.J. / Billeter, M. / Love, R.A. / Stroud, R.M. / Kuntz, I.D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1kl8.cif.gz 1kl8.cif.gz | 40 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1kl8.ent.gz pdb1kl8.ent.gz | 31.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1kl8.json.gz 1kl8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kl/1kl8 https://data.pdbj.org/pub/pdb/validation_reports/kl/1kl8 ftp://data.pdbj.org/pub/pdb/validation_reports/kl/1kl8 ftp://data.pdbj.org/pub/pdb/validation_reports/kl/1kl8 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 8005.281 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Bungarus multicinctus (many-banded krait) / References: UniProt: P60615 Bungarus multicinctus (many-banded krait) / References: UniProt: P60615 |
|---|---|
| #2: Protein/peptide | Mass: 2348.629 Da / Num. of mol.: 1 / Fragment: ALPHA-NEUROTOXIN BINDING SITE Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||||||||||||||
| NMR details | Text: THE WELL-DEFINED REGIONS IN THE COMPLEX, EXHIBITING EXTENSIVE LONG RANGE NOES, ARE: BGTX: 1-16, 22-28, 39-48, AND 54-68 ALPHA 7 19MER: 185-194. |
- Sample preparation
Sample preparation
| Details | Contents: 1.6 MM ALPHA-BUNGAROTOXIN 15N]-ALPHA 7 19MER COMPLEX MM SODIUM PHOSPHATE, PH 5. MICROMOLAR SODIUM AZIDE 50 MICROMOLAR TMSP Solvent system: 95% H2O/5% D2O |
|---|---|
| Sample conditions | pH: 5.5 / Pressure: AMBIENT / Temperature: 308.00 K |
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: DISTANCE GEOMETRY DYNAMIC SIMULATED ANNEALING / Software ordinal: 1 Details: RMS DEVIATIONS FROM IDEAL VALUES BOND LENGTH(A) 0.0039, ANGLES(DEG) 0.61, IMPROPERS(DEG) 0.5 | ||||||||||||||||||||
| NMR representative | Selection criteria: minimized average structure | ||||||||||||||||||||
| NMR ensemble | Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller




















 PDBj
PDBj


 HSQC
HSQC