[English] 日本語
 Yorodumi
Yorodumi- PDB-1hne: Structure of human neutrophil elastase in complex with a peptide ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1hne | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structure of human neutrophil elastase in complex with a peptide chloromethyl ketone inhibitor at 1.84-angstroms resolution | ||||||
 Components Components |
| ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE INHIBITOR / HYDROLASE-HYDROLASE INHIBITOR COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationleukocyte elastase / biosynthetic process of antibacterial peptides active against Gram-negative bacteria / Expression of NOTCH2NL genes / acute inflammatory response to antigenic stimulus / neutrophil-mediated killing of fungus / negative regulation of chemotaxis / positive regulation of leukocyte tethering or rolling / leukocyte migration involved in inflammatory response / response to yeast / negative regulation of chemokine production ...leukocyte elastase / biosynthetic process of antibacterial peptides active against Gram-negative bacteria / Expression of NOTCH2NL genes / acute inflammatory response to antigenic stimulus / neutrophil-mediated killing of fungus / negative regulation of chemotaxis / positive regulation of leukocyte tethering or rolling / leukocyte migration involved in inflammatory response / response to yeast / negative regulation of chemokine production / negative regulation of interleukin-8 production / Antimicrobial peptides / neutrophil-mediated killing of gram-negative bacterium / Activation of Matrix Metalloproteinases / positive regulation of MAP kinase activity / cytokine binding / Collagen degradation / extracellular matrix disassembly / Pyroptosis / pyroptotic inflammatory response / response to UV / phagocytosis / Degradation of the extracellular matrix / phagocytic vesicle / transcription repressor complex / positive regulation of smooth muscle cell proliferation / secretory granule / Regulation of Complement cascade / positive regulation of interleukin-8 production / protein catabolic process / specific granule lumen / negative regulation of inflammatory response / positive regulation of immune response / intracellular calcium ion homeostasis / azurophil granule lumen / transcription corepressor activity / peptidase activity / heparin binding / extracellular matrix / protease binding / endopeptidase activity / response to lipopolysaccharide / defense response to bacterium / serine-type endopeptidase activity / Neutrophil degranulation / cell surface / negative regulation of transcription by RNA polymerase II / Golgi apparatus / proteolysis / : / extracellular exosome / extracellular region / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.84 Å X-RAY DIFFRACTION / Resolution: 1.84 Å | ||||||
 Authors Authors | Navia, M.A. / Mckeever, B.M. / Springer, J.P. / Lin, T.-Y. / Williams, H.R. / Fluder, E.M. / Dorn, C.P. / Hoogsteen, K. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 1989 Journal: Proc.Natl.Acad.Sci.USA / Year: 1989Title: Structure of human neutrophil elastase in complex with a peptide chloromethyl ketone inhibitor at 1.84-A resolution. Authors: Navia, M.A. / McKeever, B.M. / Springer, J.P. / Lin, T.Y. / Williams, H.R. / Fluder, E.M. / Dorn, C.P. / Hoogsteen, K. #1:  Journal: J.Biol.Chem. / Year: 1987 Journal: J.Biol.Chem. / Year: 1987Title: Crystallization of Human Neutrophil Elastase Authors: Williams, H.R. / Lin, T.-Y. / Navia, M.A. / Springer, J.P. / Mckeever, B.M. / Hoogsteen, K. / Dornjunior, C.P. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1hne.cif.gz 1hne.cif.gz | 61.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1hne.ent.gz pdb1hne.ent.gz | 43.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1hne.json.gz 1hne.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/hn/1hne https://data.pdbj.org/pub/pdb/validation_reports/hn/1hne ftp://data.pdbj.org/pub/pdb/validation_reports/hn/1hne ftp://data.pdbj.org/pub/pdb/validation_reports/hn/1hne | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 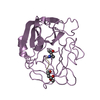
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 23319.967 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / References: UniProt: P08246, leukocyte elastase Homo sapiens (human) / References: UniProt: P08246, leukocyte elastase |
|---|---|
| #2: Protein/peptide | |
| #3: Water | ChemComp-HOH / |
| Compound details | THE UNBOUND FORM OF THE INHIBITOR (CHAIN I) IS METHOXYSUCCINYL-ALA-ALA-PRO-ALA-CHLOROMETHYLKETONE. ...THE UNBOUND FORM OF THE INHIBITOR (CHAIN I) IS METHOXYSUC |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.39 Å3/Da / Density % sol: 48.55 % | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Temperature: 20 ℃ / pH: 5 / Method: vapor diffusion, hanging drop / Details: referred to J.Biol.Chem. 262.17178-17181 1987 | ||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | Highest resolution: 1.84 Å |
| Reflection | *PLUS Highest resolution: 1.84 Å / Rmerge(I) obs: 0.165 |
- Processing
Processing
| Software | Name: PROLSQ / Classification: refinement | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor obs: 0.164 / Highest resolution: 1.84 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 1.84 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Lowest resolution: 8 Å / Num. reflection obs: 16828 / Highest resolution: 1.84 Å / Rfactor obs: 0.164 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj