[English] 日本語
 Yorodumi
Yorodumi- PDB-1csk: THE CRYSTAL STRUCTURE OF HUMAN CSKSH3: STRUCTURAL DIVERSITY NEAR ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1csk | ||||||
|---|---|---|---|---|---|---|---|
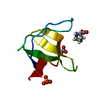
| Title | THE CRYSTAL STRUCTURE OF HUMAN CSKSH3: STRUCTURAL DIVERSITY NEAR THE RT-SRC AND N-SRC LOOP | ||||||
 Components Components | C-SRC SH3 DOMAIN | ||||||
 Keywords Keywords | PHOSPHOTRANSFERASE | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of Fc receptor mediated stimulatory signaling pathway / negative regulation of Golgi to plasma membrane protein transport / adherens junction organization / proline-rich region binding / negative regulation of low-density lipoprotein particle clearance / negative regulation of bone resorption / cellular response to peptide hormone stimulus / negative regulation of phagocytosis / negative regulation of T cell activation / Phosphorylation of CD3 and TCR zeta chains ...regulation of Fc receptor mediated stimulatory signaling pathway / negative regulation of Golgi to plasma membrane protein transport / adherens junction organization / proline-rich region binding / negative regulation of low-density lipoprotein particle clearance / negative regulation of bone resorption / cellular response to peptide hormone stimulus / negative regulation of phagocytosis / negative regulation of T cell activation / Phosphorylation of CD3 and TCR zeta chains / protein kinase A catalytic subunit binding / oligodendrocyte differentiation / negative regulation of interleukin-6 production / RHOH GTPase cycle / Co-inhibition by PD-1 / negative regulation of T cell receptor signaling pathway / GAB1 signalosome / T cell costimulation / Integrin signaling / Negative regulation of FLT3 / protein tyrosine kinase binding / non-specific protein-tyrosine kinase / non-membrane spanning protein tyrosine kinase activity / Signaling by high-kinase activity BRAF mutants / MAP2K and MAPK activation / negative regulation of ERK1 and ERK2 cascade / Signaling by RAF1 mutants / Signaling by moderate kinase activity BRAF mutants / Paradoxical activation of RAF signaling by kinase inactive BRAF / Signaling downstream of RAS mutants / cell-cell junction / Signaling by BRAF and RAF1 fusions / T cell receptor signaling pathway / protein tyrosine kinase activity / protein phosphatase binding / protein phosphorylation / adaptive immune response / intracellular signal transduction / negative regulation of cell population proliferation / extracellular exosome / ATP binding / metal ion binding / identical protein binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.5 Å X-RAY DIFFRACTION / Resolution: 2.5 Å | ||||||
 Authors Authors | Mathieu, M. / Wierenga, R.K. | ||||||
 Citation Citation |  Journal: FEBS Lett. / Year: 1994 Journal: FEBS Lett. / Year: 1994Title: The crystal structure of human CskSH3: structural diversity near the RT-Src and n-Src loop. Authors: Borchert, T.V. / Mathieu, M. / Zeelen, J.P. / Courtneidge, S.A. / Wierenga, R.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1csk.cif.gz 1csk.cif.gz | 54.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1csk.ent.gz pdb1csk.ent.gz | 41 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1csk.json.gz 1csk.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/cs/1csk https://data.pdbj.org/pub/pdb/validation_reports/cs/1csk ftp://data.pdbj.org/pub/pdb/validation_reports/cs/1csk ftp://data.pdbj.org/pub/pdb/validation_reports/cs/1csk | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | THE FOUR MOLECULES ARE PACKED IN THE CELL AS A DIMER OF DIMERS, ONE DIMER BEING FORMED OF MOLECULES D AND A, AND THE OTHER OF MOLECULES B AND C. |
- Components
Components
| #1: Protein | Mass: 7851.946 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / References: UniProt: P41240, EC: 2.7.1.112 Homo sapiens (human) / References: UniProt: P41240, EC: 2.7.1.112#2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.41 Å3/Da / Density % sol: 48.98 % | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Method: microdialysis | ||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Reflection | *PLUS Highest resolution: 2.5 Å / Lowest resolution: 25 Å / Num. obs: 10807 / % possible obs: 94.8 % / Num. measured all: 60252 / Rmerge(I) obs: 0.083 |
|---|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.5→8 Å / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.5→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Num. reflection all: 9913 / Rfactor Rfree: 0.278 / Rfactor Rwork: 0.224 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: x_angle_d / Dev ideal: 1.9 |
 Movie
Movie Controller
Controller







 PDBj
PDBj














