+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-7480 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of mouse RAG1/2 HFC complex containing partial HMGB1 linker(3.9 A) | |||||||||
 マップデータ マップデータ | Cryo-EM structure of mouse RAG1/2 HFC complex | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | V(D)J recombination / RAG1/2 / RSS / Immunity / RECOMBINATION / RECOMBINATION-DNA complex | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報: / regulation of tolerance induction / calcium-dependent protein kinase regulator activity / positive regulation of myeloid progenitor cell differentiation / regulation of T cell mediated immune response to tumor cell / positive regulation of mismatch repair / negative regulation of apoptotic cell clearance / plasmacytoid dendritic cell activation / negative regulation of RNA polymerase II transcription preinitiation complex assembly / myeloid dendritic cell activation ...: / regulation of tolerance induction / calcium-dependent protein kinase regulator activity / positive regulation of myeloid progenitor cell differentiation / regulation of T cell mediated immune response to tumor cell / positive regulation of mismatch repair / negative regulation of apoptotic cell clearance / plasmacytoid dendritic cell activation / negative regulation of RNA polymerase II transcription preinitiation complex assembly / myeloid dendritic cell activation / T-helper 1 cell activation / T-helper 1 cell differentiation / mature B cell differentiation involved in immune response / positive regulation of myeloid cell differentiation / positive regulation of toll-like receptor 2 signaling pathway / positive regulation of dendritic cell differentiation / C-X-C chemokine binding / negative regulation of CD4-positive, alpha-beta T cell differentiation / positive regulation of toll-like receptor 9 signaling pathway / B cell homeostatic proliferation / neutrophil clearance / negative regulation of T cell differentiation in thymus / positive regulation of glycogen catabolic process / DN2 thymocyte differentiation / pre-B cell allelic exclusion / DNA geometric change / positive regulation of toll-like receptor 4 signaling pathway / endothelial cell chemotaxis / RAGE receptor binding / positive regulation of organ growth / eye development / positive regulation of interleukin-1 production / Regulation of TLR by endogenous ligand / bubble DNA binding / regulation of behavioral fear response / V(D)J recombination / negative regulation of T cell apoptotic process / myeloid cell differentiation / alphav-beta3 integrin-HMGB1 complex / Apoptosis induced DNA fragmentation / myeloid progenitor cell differentiation / inflammatory response to antigenic stimulus / phosphatidylinositol-3,4-bisphosphate binding / macrophage activation involved in immune response / negative regulation of thymocyte apoptotic process / histone H3K4me3 reader activity / positive regulation of monocyte chemotaxis / MyD88 deficiency (TLR2/4) / positive regulation of monocyte chemotactic protein-1 production / regulation of nucleotide-excision repair / cellular response to interleukin-7 / endothelial cell proliferation / phosphatidylinositol-3,5-bisphosphate binding / positive regulation of chemokine (C-X-C motif) ligand 2 production / glycogen catabolic process / apoptotic cell clearance / positive regulation of vascular endothelial cell proliferation / IRAK4 deficiency (TLR2/4) / supercoiled DNA binding / regulation of T cell differentiation / dendritic cell chemotaxis / MyD88:MAL(TIRAP) cascade initiated on plasma membrane / positive regulation of DNA binding / DNA binding, bending / organ growth / positive regulation of T cell differentiation / T cell lineage commitment / positive regulation of wound healing / phosphatidylserine binding / B cell lineage commitment / positive regulation of sprouting angiogenesis / chemoattractant activity / endoplasmic reticulum-Golgi intermediate compartment / phosphatidylinositol-3,4,5-trisphosphate binding / TRAF6 mediated NF-kB activation / T cell homeostasis / positive regulation of activated T cell proliferation / negative regulation of type II interferon production / DNA topological change / Advanced glycosylation endproduct receptor signaling / positive regulation of interferon-alpha production / positive regulation of interleukin-10 production / negative regulation of blood vessel endothelial cell migration / T cell differentiation / positive regulation of blood vessel endothelial cell migration / Pyroptosis / protein kinase activator activity / protein autoubiquitination / four-way junction DNA binding / condensed chromosome / DNA polymerase binding / phosphatidylinositol-4,5-bisphosphate binding / transcription repressor complex / positive regulation of interleukin-12 production / positive regulation of autophagy / phosphatidylinositol binding / activation of innate immune response / positive regulation of interferon-beta production / lung development / response to glucocorticoid 類似検索 - 分子機能 | |||||||||
| 生物種 |   Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.9 Å | |||||||||
 データ登録者 データ登録者 | Chen X / Kim M | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Mol Cell / 年: 2018 ジャーナル: Mol Cell / 年: 2018タイトル: Cracking the DNA Code for V(D)J Recombination. 著者: Min-Sung Kim / Watchalee Chuenchor / Xuemin Chen / Yanxiang Cui / Xing Zhang / Z Hong Zhou / Martin Gellert / Wei Yang /    要旨: To initiate V(D)J recombination for generating the adaptive immune response of vertebrates, RAG1/2 recombinase cleaves DNA at a pair of recombination signal sequences, the 12- and 23-RSS. We have ...To initiate V(D)J recombination for generating the adaptive immune response of vertebrates, RAG1/2 recombinase cleaves DNA at a pair of recombination signal sequences, the 12- and 23-RSS. We have determined crystal and cryo-EM structures of RAG1/2 with DNA in the pre-reaction and hairpin-forming complexes up to 2.75 Å resolution. Both protein and DNA exhibit structural plasticity and undergo dramatic conformational changes. Coding-flank DNAs extensively rotate, shift, and deform for nicking and hairpin formation. Two intertwined RAG1 subunits crisscross four times between the asymmetric pair of severely bent 12/23-RSS DNAs. Location-sensitive bending of 60° and 150° in 12- and 23-RSS spacers, respectively, must occur for RAG1/2 to capture the nonamers and pair the heptamers for symmetric double-strand breakage. DNA pairing is thus sequence-context dependent and structure specific, which partly explains the "beyond 12/23" restriction. Finally, catalysis in crystallo reveals the process of DNA hairpin formation and its stabilization by interleaved base stacking. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_7480.map.gz emd_7480.map.gz | 8.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-7480-v30.xml emd-7480-v30.xml emd-7480.xml emd-7480.xml | 25 KB 25 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_7480.png emd_7480.png | 169.9 KB | ||
| Filedesc metadata |  emd-7480.cif.gz emd-7480.cif.gz | 7.8 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7480 http://ftp.pdbj.org/pub/emdb/structures/EMD-7480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7480 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_7480_validation.pdf.gz emd_7480_validation.pdf.gz | 379.2 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_7480_full_validation.pdf.gz emd_7480_full_validation.pdf.gz | 378.8 KB | 表示 | |
| XML形式データ |  emd_7480_validation.xml.gz emd_7480_validation.xml.gz | 6.4 KB | 表示 | |
| CIF形式データ |  emd_7480_validation.cif.gz emd_7480_validation.cif.gz | 7.4 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7480 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7480 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7480 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7480 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  6cijMC  7470C  5zdzC  5ze0C  5ze1C  5ze2C  6cg0C  6cikC  6cilC  6cimC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_7480.map.gz / 形式: CCP4 / 大きさ: 91.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_7480.map.gz / 形式: CCP4 / 大きさ: 91.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Cryo-EM structure of mouse RAG1/2 HFC complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
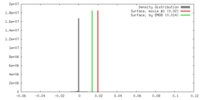
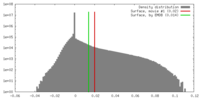
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : RAG1/2 in complex with nicked DNAs
+超分子 #1: RAG1/2 in complex with nicked DNAs
+分子 #1: V(D)J recombination-activating protein 1
+分子 #6: High mobility group protein B1
+分子 #8: V(D)J recombination-activating protein 2
+分子 #2: DNA (46-MER)
+分子 #3: DNA (5'-D(*GP*AP*TP*CP*TP*GP*GP*CP*CP*TP*GP*TP*CP*TP*TP*A)-3')
+分子 #4: DNA (5'-D(P*CP*TP*GP*GP*AP*TP*CP*TP*GP*GP*CP*CP*TP*GP*TP*CP*TP*TP...
+分子 #5: DNA (60-MER)
+分子 #7: DNA (30-MER)
+分子 #9: DNA (41-MER)
+分子 #10: ZINC ION
+分子 #11: CALCIUM ION
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.3 mg/mL |
|---|---|
| 緩衝液 | pH: 7.4 |
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 平均電子線量: 57.6 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 精密化 | プロトコル: RIGID BODY FIT / 当てはまり具合の基準: Correlation Coefficient |
|---|---|
| 得られたモデル |  PDB-6cij: |
 ムービー
ムービー コントローラー
コントローラー



































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)