+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6875 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of atOSCA3.1 channel | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | osca / TMEM63 / ion channel / mechanosensitive / membrane protein / osmosensing / METAL TRANSPORT | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationplasmodesma / mechanosensitive monoatomic ion channel activity / plant-type vacuole / chloroplast envelope / calcium-activated cation channel activity / mRNA binding / nucleus / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
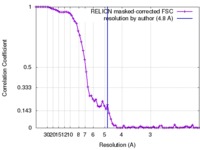
| Method | single particle reconstruction / cryo EM / Resolution: 4.8 Å | ||||||||||||
 Authors Authors | Chen L / Zhang M | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2018 Journal: Nat Struct Mol Biol / Year: 2018Title: Structure of the mechanosensitive OSCA channels. Authors: Mingfeng Zhang / Dali Wang / Yunlu Kang / Jing-Xiang Wu / Fuqiang Yao / Chengfang Pan / Zhiqiang Yan / Chen Song / Lei Chen /  Abstract: Mechanosensitive ion channels convert mechanical stimuli into a flow of ions. These channels are widely distributed from bacteria to higher plants and humans, and are involved in many crucial ...Mechanosensitive ion channels convert mechanical stimuli into a flow of ions. These channels are widely distributed from bacteria to higher plants and humans, and are involved in many crucial physiological processes. Here we show that two members of the OSCA protein family in Arabidopsis thaliana, namely AtOSCA1.1 and AtOSCA3.1, belong to a new class of mechanosensitive ion channels. We solve the structure of the AtOSCA1.1 channel at 3.5-Å resolution and AtOSCA3.1 at 4.8-Å resolution by cryo-electron microscopy. OSCA channels are symmetric dimers that are mediated by cytosolic inter-subunit interactions. Strikingly, they have structural similarity to the mammalian TMEM16 family proteins. Our structural analysis accompanied with electrophysiological studies identifies the ion permeation pathway within each subunit and suggests a conformational change model for activation. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6875.map.gz emd_6875.map.gz | 4.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6875-v30.xml emd-6875-v30.xml emd-6875.xml emd-6875.xml | 15.6 KB 15.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6875_fsc.xml emd_6875_fsc.xml | 7 KB | Display |  FSC data file FSC data file |
| Images |  emd_6875.png emd_6875.png | 117.8 KB | ||
| Filedesc metadata |  emd-6875.cif.gz emd-6875.cif.gz | 5.6 KB | ||
| Others |  emd_6875_half_map_1.map.gz emd_6875_half_map_1.map.gz emd_6875_half_map_2.map.gz emd_6875_half_map_2.map.gz | 22.5 MB 22.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6875 http://ftp.pdbj.org/pub/emdb/structures/EMD-6875 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6875 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6875 | HTTPS FTP |
-Related structure data
| Related structure data |  5z1fMC  6822C  6jpfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6875.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6875.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.055 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #1
| File | emd_6875_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
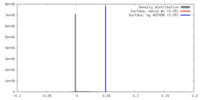
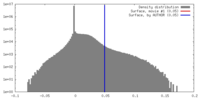
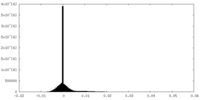
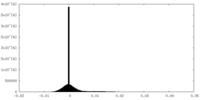
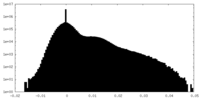
| Density Histograms |
-Half map: #2
| File | emd_6875_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : atOSCA3.1
| Entire | Name: atOSCA3.1 |
|---|---|
| Components |
|
-Supramolecule #1: atOSCA3.1
| Supramolecule | Name: atOSCA3.1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 175 KDa |
-Macromolecule #1: CSC1-like protein ERD4
| Macromolecule | Name: CSC1-like protein ERD4 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 82.017727 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEFGSFLVSL GTSFVIFVIL MLLFTWLSRK SGNAPIYYPN RILKGLEPWE GTSLTRNPFA WMREALTSSE QDVVNLSGVD TAVHFVFLS TVLGIFACSS LLLLPTLLPL AATDNNIKNT KNATDTTSKG TFSQLDNLSM ANITKKSSRL WAFLGAVYWI S LVTYFFLW ...String: MEFGSFLVSL GTSFVIFVIL MLLFTWLSRK SGNAPIYYPN RILKGLEPWE GTSLTRNPFA WMREALTSSE QDVVNLSGVD TAVHFVFLS TVLGIFACSS LLLLPTLLPL AATDNNIKNT KNATDTTSKG TFSQLDNLSM ANITKKSSRL WAFLGAVYWI S LVTYFFLW KAYKHVSSLR AQALMSADVK PEQFAILVRD MPAPPDGQTQ KEFIDSYFRE IYPETFYRSL VATENSKVNK IW EKLEGYK KKLARAEAIL AATNNRPTNK TGFCGLVGKQ VDSIEYYTEL INESVAKLET EQKAVLAEKQ QTAAVVFFTT RVA AASAAQ SLHCQMVDKW TVTEAPEPRQ LLWQNLNIKL FSRIIRQYFI YFFVAVTILF YMIPIAFVSA ITTLKNLQRI IPFI KPVVE ITAIRTVLES FLPQIALIVF LAMLPKLLLF LSKAEGIPSQ SHAIRAASGK YFYFSVFNVF IGVTLAGTLF NTVKD IAKN PKLDMIINLL ATSLPKSATF FLTYVALKFF IGYGLELSRI IPLIIFHLKK KYLCKTEAEV KEAWYPGDLS YATRVP GDM LILTITFCYS VIAPLILIFG ITYFGLGWLV LRNQALKVYV PSYESYGRMW PHIHQRILAA LFLFQVVMFG YLGAKTF FY TALVIPLIIT SLIFGYVCRQ KFYGGFEHTA LEVACRELKQ SPDLEEIFRA YIPHSLSSHK PEEHEFKGAM SRYQDFNA I AGV UniProtKB: Hyperosmolality-gated Ca2+ permeable channel 3.1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-5z1f: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)