[English] 日本語
 Yorodumi
Yorodumi- EMDB-5741: Single-particle Electron Microscopy Structure of the TRAPPIII Complex -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5741 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Single-particle Electron Microscopy Structure of the TRAPPIII Complex | |||||||||
 Map data Map data | Reconstruction of yeast TRAPPIII complex, filtered to 22 Angstrom resolution | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | autophagy | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 22.0 Å | |||||||||
 Authors Authors | Tan D / Cai Y / Wang J / Zhang J / Menon S / Chou H-T / Ferro-Novick S / Reinisch KM / Walz T | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2013 Journal: Proc Natl Acad Sci U S A / Year: 2013Title: The EM structure of the TRAPPIII complex leads to the identification of a requirement for COPII vesicles on the macroautophagy pathway. Authors: Dongyan Tan / Yiying Cai / Juan Wang / Jinzhong Zhang / Shekar Menon / Hui-Ting Chou / Susan Ferro-Novick / Karin M Reinisch / Thomas Walz /  Abstract: The transport protein particle (TRAPP) III complex, comprising the TRAPPI complex and additional subunit Trs85, is an autophagy-specific guanine nucleotide exchange factor for the Rab GTPase Ypt1 ...The transport protein particle (TRAPP) III complex, comprising the TRAPPI complex and additional subunit Trs85, is an autophagy-specific guanine nucleotide exchange factor for the Rab GTPase Ypt1 that is recruited to the phagophore assembly site when macroautophagy is induced. We present the single-particle electron microscopy structure of TRAPPIII, which reveals that the dome-shaped Trs85 subunit associates primarily with the Trs20 subunit of TRAPPI. We further demonstrate that TRAPPIII binds the coat protein complex (COP) II coat subunit Sec23. The COPII coat facilitates the budding and targeting of ER-derived vesicles with their acceptor compartment. We provide evidence that COPII-coated vesicles and the ER-Golgi fusion machinery are needed for macroautophagy. Our results imply that TRAPPIII binds to COPII vesicles at the phagophore assembly site and that COPII vesicles may provide one of the membrane sources used in autophagosome formation. These events are conserved in yeast to mammals. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5741.map.gz emd_5741.map.gz | 1.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5741-v30.xml emd-5741-v30.xml emd-5741.xml emd-5741.xml | 10.6 KB 10.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5741_1.tif emd_5741_1.tif | 119.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5741 http://ftp.pdbj.org/pub/emdb/structures/EMD-5741 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5741 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5741 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5741.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5741.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of yeast TRAPPIII complex, filtered to 22 Angstrom resolution | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.48 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
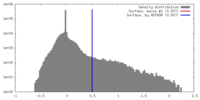
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Recombinant TRAPPIII complex
| Entire | Name: Recombinant TRAPPIII complex |
|---|---|
| Components |
|
-Supramolecule #1000: Recombinant TRAPPIII complex
| Supramolecule | Name: Recombinant TRAPPIII complex / type: sample / ID: 1000 / Oligomeric state: oligomer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 254 KDa / Theoretical: 254 KDa |
-Macromolecule #1: transport protein particle III
| Macromolecule | Name: transport protein particle III / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 254 KDa / Theoretical: 254 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 20mM Tris-HCl, 300 mM NaCl, 1mM DTT |
| Staining | Type: NEGATIVE Details: Grids with adsorbed protein stained with 0.75% uranyl format for 20 seconds |
| Grid | Details: 200 mesh copper grids with thin carbon support, glow discharged |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Details | low dose setting |
| Date | Dec 20, 2010 |
| Image recording | Category: FILM / Film or detector model: GENERIC IMAGE PLATES / Digitization - Scanner: OTHER / Digitization - Sampling interval: 15 µm / Number real images: 133 / Average electron dose: 10 e/Å2 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Calibrated magnification: 67000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 67000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC / Tilt angle max: 60 |
- Image processing
Image processing
| Details | Random Conical Tilt reconstruction using Spider scripts |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 22.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Spider Details: Final maps were calculated from a dataset of two class-averages Number images used: 1120 |
| Final two d classification | Number classes: 20 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)