[English] 日本語
 Yorodumi
Yorodumi- EMDB-5127: Three-dimensional EM structure of an intact activator-dependent t... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5127 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Three-dimensional EM structure of an intact activator-dependent transcription initiation complex | |||||||||
 Map data Map data | E. coli Class I transcription activation complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transcription / initiation / Class I / activator / RNA polymerase / holoenzyme / sigma70 / open complex / CAP / CRP / cAMP-dependent / DNA / prokaryotic | |||||||||
| Function / homology |  Function and homology information Function and homology informationcarbon catabolite repression of transcription / sigma factor antagonist complex / DNA binding, bending / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / minor groove of adenine-thymine-rich DNA binding / regulation of DNA-templated transcription initiation / sigma factor activity / bacterial-type flagellum assembly ...carbon catabolite repression of transcription / sigma factor antagonist complex / DNA binding, bending / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / minor groove of adenine-thymine-rich DNA binding / regulation of DNA-templated transcription initiation / sigma factor activity / bacterial-type flagellum assembly / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / bacterial-type flagellum-dependent cell motility / nitrate assimilation / cAMP binding / regulation of DNA-templated transcription elongation / transcription elongation factor complex / DNA-directed RNA polymerase complex / transcription antitermination / DNA-templated transcription initiation / cell motility / protein-DNA complex / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / response to heat / protein-containing complex assembly / sequence-specific DNA binding / intracellular iron ion homeostasis / protein dimerization activity / transcription cis-regulatory region binding / DNA-binding transcription factor activity / response to antibiotic / negative regulation of DNA-templated transcription / regulation of DNA-templated transcription / DNA-templated transcription / positive regulation of DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / membrane / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 19.8 Å | |||||||||
 Authors Authors | Hudson BP / Quispe J / Lara S / Kim Y / Berman HM / Arnold E / Ebright RH / Lawson CL | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2009 Journal: Proc Natl Acad Sci U S A / Year: 2009Title: Three-dimensional EM structure of an intact activator-dependent transcription initiation complex. Authors: Brian P Hudson / Joel Quispe / Samuel Lara-González / Younggyu Kim / Helen M Berman / Eddy Arnold / Richard H Ebright / Catherine L Lawson /  Abstract: We present the experimentally determined 3D structure of an intact activator-dependent transcription initiation complex comprising the Escherichia coli catabolite activator protein (CAP), RNA ...We present the experimentally determined 3D structure of an intact activator-dependent transcription initiation complex comprising the Escherichia coli catabolite activator protein (CAP), RNA polymerase holoenzyme (RNAP), and a DNA fragment containing positions -78 to +20 of a Class I CAP-dependent promoter with a CAP site at position -61.5 and a premelted transcription bubble. A 20-A electron microscopy reconstruction was obtained by iterative projection-based matching of single particles visualized in carbon-sandwich negative stain and was fitted using atomic coordinate sets for CAP, RNAP, and DNA. The structure defines the organization of a Class I CAP-RNAP-promoter complex and supports previously proposed interactions of CAP with RNAP alpha subunit C-terminal domain (alphaCTD), interactions of alphaCTD with sigma(70) region 4, interactions of CAP and RNAP with promoter DNA, and phased-DNA-bend-dependent partial wrapping of DNA around the complex. The structure also reveals the positions and shapes of species-specific domains within the RNAP beta', beta, and sigma(70) subunits. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5127.map.gz emd_5127.map.gz | 1.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5127-v30.xml emd-5127-v30.xml emd-5127.xml emd-5127.xml | 18.2 KB 18.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5127_1.tif emd_5127_1.tif | 764.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5127 http://ftp.pdbj.org/pub/emdb/structures/EMD-5127 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5127 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5127 | HTTPS FTP |
-Related structure data
| Related structure data |  3iydMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5127.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5127.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E. coli Class I transcription activation complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.64 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
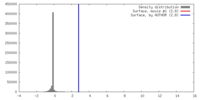
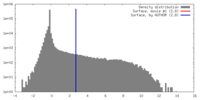
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : E. coli RNA polymerase holoenzyme (sigma70) and E. coli catabolit...
| Entire | Name: E. coli RNA polymerase holoenzyme (sigma70) and E. coli catabolite activator protein (CAP) bound to 98-mer DNA containing the lac promoter and engineered open transcription bubble |
|---|---|
| Components |
|
-Supramolecule #1000: E. coli RNA polymerase holoenzyme (sigma70) and E. coli catabolit...
| Supramolecule | Name: E. coli RNA polymerase holoenzyme (sigma70) and E. coli catabolite activator protein (CAP) bound to 98-mer DNA containing the lac promoter and engineered open transcription bubble type: sample / ID: 1000 / Details: Complex formation was verified by gel shift Oligomeric state: One molecule of RNAP (containing six subunits) and one CAP homodimer bound to a DNA duplex) Number unique components: 3 |
|---|---|
| Molecular weight | Theoretical: 570 KDa |
-Macromolecule #1: Catabolite Activator Protein
| Macromolecule | Name: Catabolite Activator Protein / type: protein_or_peptide / ID: 1 / Name.synonym: CAP / Number of copies: 1 / Oligomeric state: homodimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 50 KDa |
| Recombinant expression | Organism:  |
| Sequence | GO: regulation of DNA-templated transcription / InterPro: INTERPRO: IPR001808 |
-Macromolecule #3: RNA polymerase holoenzyme (sigma70)
| Macromolecule | Name: RNA polymerase holoenzyme (sigma70) / type: protein_or_peptide / ID: 3 / Name.synonym: RNAP Details: Six subunits include alphaI (RpoA), alphaII (RpoA), beta (RpoB), beta prime (RpoC) with C-terminal 6His-tag, omega (RpoZ), and sigma70. Number of copies: 1 / Oligomeric state: heterohexamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 460 KDa |
| Recombinant expression | Organism:  |
| Sequence | GO: DNA-templated transcription |
-Macromolecule #2: lac(ICAP)UP-UV5-bubble
| Macromolecule | Name: lac(ICAP)UP-UV5-bubble / type: dna / ID: 2 / Name.synonym: lac(ICAP)UP-UV5-bubble Details: An engineered 98mer duplex based on positions -78 to 20 of the Class I CAP-dependent promoter lac but containing consensus -10 sequence and consensus binding sites for CAP and RNAP alpha-CTD. ...Details: An engineered 98mer duplex based on positions -78 to 20 of the Class I CAP-dependent promoter lac but containing consensus -10 sequence and consensus binding sites for CAP and RNAP alpha-CTD. Positions -11 to 2 are non-complementary to create an artificial transcription bubble. Top strand 5'-CGCAATAAATGTGATCTAGATCACATTTTAGGCAAAAAAGGCTTTACACTTTATGCTTCCGGCTCGTATAATCGCACCTTATGTGAGCGGATAACAAG-3' Bottom strand 5'-CTTGTTATCCGCTCACAATTCCACACTAATAACGAGCCGGAAGCATAAAGTGTAAAGCCTTTTTTGCCTAAAATGTGATCTAGATCACATTTATTGCG-3' Classification: DNA / Structure: OTHER / Synthetic?: Yes |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Experimental: 60 KDa / Theoretical: 60 KDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6.18 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 25mM HEPES, 100mM KCl, 10mM MgCl2, 1mM DTT, 0.2mM cAMP |
| Staining | Type: NEGATIVE Details: Sample and 2% uranyl formate stain were applied to the grid in rapid succession, with the last drop of stain remaining on the sample for 1 minute. The grid was then submerged in stain and ...Details: Sample and 2% uranyl formate stain were applied to the grid in rapid succession, with the last drop of stain remaining on the sample for 1 minute. The grid was then submerged in stain and brought up under thin carbon to form an upper sandwich layer. The grid was then blotted and dried for 10 minutes. |
| Grid | Details: 400-mesh copper 2.0x0.5 hole pattern C-flat grid covered with thin layer of continuous carbon |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Average: 293 K |
| Details | 15 um pixel size on detector |
| Date | Nov 4, 2008 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F415 (4k x 4k) / Number real images: 349 / Average electron dose: 16 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: standard side-entry room-temperature stage / Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | 32816 particles were automatically selected by the Appion DoGpicker initially |
|---|---|
| CTF correction | Details: ACE |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 19.8 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN, SPIDER Details: EMAN interleaved with SPIDER correspondence analysis Number images used: 14097 |
| Final two d classification | Number classes: 280 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: Chimera, Yup.scx |
| Details | Protocol: rigid body, Yup.scx simulated annealing. A complete ternary complex model was generated using multiple PDB entries, with a homology modelling step for RNAP. The model was regularized with PHENIX and the refined against the EM map with Yup.scx using default parameters. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: map-derived potential energy |
| Output model |  PDB-3iyd: |
-Atomic model buiding 2
| Initial model | PDB ID: |
|---|---|
| Software | Name: Chimera, Yup.scx |
| Details | Protocol: rigid body fit followed by Yup.scx simulated annealing. A complete ternary complex model was generated using multiple PDB entries, with a homology modelling step for RNAP. The model was regularized with PHENIX and the refined against the EM map with Yup.scx using default parameters. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: map-derived potential energy |
| Output model |  PDB-3iyd: |
-Atomic model buiding 3
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name: Chimera, Yup.scx |
| Details | Protocol: manual fit followed by Yup.scx simulated annealing. A complete ternary complex model was generated using multiple PDB entries, with a homology modelling step for RNAP. The model was regularized with PHENIX and the refined against the EM map with Yup.scx using default parameters. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: map-derived potential energy |
| Output model |  PDB-3iyd: |
-Atomic model buiding 4
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name: Chimera, Yup.scx |
| Details | Protocol: rigid body fit followed by Yup.scx simulated annealing. A complete ternary complex model was generated using multiple PDB entries, with a homology modelling step for RNAP. The model was regularized with PHENIX and the refined against the EM map with Yup.scx using default parameters. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: map-derived potential energy |
| Output model |  PDB-3iyd: |
-Atomic model buiding 5
| Initial model | PDB ID: |
|---|---|
| Software | Name: Chimera, Modeller, Yup.scx |
| Details | Protocol: rigid body fit followed by Yup.scx simulated annealing. A complete ternary complex model was generated using multiple PDB entries, with a homology modelling step for RNAP. The model was regularized with PHENIX and the refined against the EM map with Yup.scx using default parameters. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: map-derived potential energy |
| Output model |  PDB-3iyd: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)