[English] 日本語
 Yorodumi
Yorodumi- EMDB-4664: Influenza A virus (A/NT/60/1968) polymerase dimer of Hetermotrime... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4664 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Influenza A virus (A/NT/60/1968) polymerase dimer of Hetermotrimer in complex with 3'5' cRNA promoter | ||||||||||||
 Map data Map data | None | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationcap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / virion component / endonuclease activity / Hydrolases; Acting on ester bonds / host cell cytoplasm ...cap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / virion component / endonuclease activity / Hydrolases; Acting on ester bonds / host cell cytoplasm / symbiont-mediated suppression of host gene expression / viral translational frameshifting / RNA-directed RNA polymerase / nucleotide binding / viral RNA genome replication / hydrolase activity / RNA-directed RNA polymerase activity / DNA-templated transcription / host cell nucleus / RNA binding / metal ion binding Similarity search - Function | ||||||||||||
| Biological species |  Influenza A virus (A/nt/60/1968(H3N2)) Influenza A virus (A/nt/60/1968(H3N2)) | ||||||||||||
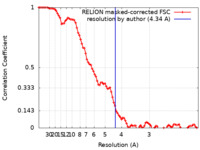
| Method | single particle reconstruction / cryo EM / Resolution: 4.34 Å | ||||||||||||
 Authors Authors | Carrique L / Keown JR | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Structures of influenza A virus RNA polymerase offer insight into viral genome replication. Authors: Haitian Fan / Alexander P Walker / Loïc Carrique / Jeremy R Keown / Itziar Serna Martin / Dimple Karia / Jane Sharps / Narin Hengrung / Els Pardon / Jan Steyaert / Jonathan M Grimes / Ervin Fodor /    Abstract: Influenza A viruses are responsible for seasonal epidemics, and pandemics can arise from the transmission of novel zoonotic influenza A viruses to humans. Influenza A viruses contain a segmented ...Influenza A viruses are responsible for seasonal epidemics, and pandemics can arise from the transmission of novel zoonotic influenza A viruses to humans. Influenza A viruses contain a segmented negative-sense RNA genome, which is transcribed and replicated by the viral-RNA-dependent RNA polymerase (FluPol) composed of PB1, PB2 and PA subunits. Although the high-resolution crystal structure of FluPol of bat influenza A virus has previously been reported, there are no complete structures available for human and avian FluPol. Furthermore, the molecular mechanisms of genomic viral RNA (vRNA) replication-which proceeds through a complementary RNA (cRNA) replicative intermediate, and requires oligomerization of the polymerase-remain largely unknown. Here, using crystallography and cryo-electron microscopy, we determine the structures of FluPol from human influenza A/NT/60/1968 (H3N2) and avian influenza A/duck/Fujian/01/2002 (H5N1) viruses at a resolution of 3.0-4.3 Å, in the presence or absence of a cRNA or vRNA template. In solution, FluPol forms dimers of heterotrimers through the C-terminal domain of the PA subunit, the thumb subdomain of PB1 and the N1 subdomain of PB2. The cryo-electron microscopy structure of monomeric FluPol bound to the cRNA template reveals a binding site for the 3' cRNA at the dimer interface. We use a combination of cell-based and in vitro assays to show that the interface of the FluPol dimer is required for vRNA synthesis during replication of the viral genome. We also show that a nanobody (a single-domain antibody) that interferes with FluPol dimerization inhibits the synthesis of vRNA and, consequently, inhibits virus replication in infected cells. Our study provides high-resolution structures of medically relevant FluPol, as well as insights into the replication mechanisms of the viral RNA genome. In addition, our work identifies sites in FluPol that could be targeted in the development of antiviral drugs. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4664.map.gz emd_4664.map.gz | 37.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4664-v30.xml emd-4664-v30.xml emd-4664.xml emd-4664.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4664_fsc.xml emd_4664_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  emd_4664.png emd_4664.png | 70.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4664 http://ftp.pdbj.org/pub/emdb/structures/EMD-4664 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4664 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4664 | HTTPS FTP |
-Related structure data
| Related structure data |  4660C  4661C  4663C  4666C  4986C  6qnwC  6qpfC  6qpgC  6qwlC  6qx3C  6qx8C  6qxeC  6rr7C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4664.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4664.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.043 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Influenza A virus (A/NT/60/1968) polymerase dimer of heterotrimer...
| Entire | Name: Influenza A virus (A/NT/60/1968) polymerase dimer of heterotrimer in complex with 5' cRNA promoter |
|---|---|
| Components |
|
-Supramolecule #1: Influenza A virus (A/NT/60/1968) polymerase dimer of heterotrimer...
| Supramolecule | Name: Influenza A virus (A/NT/60/1968) polymerase dimer of heterotrimer in complex with 5' cRNA promoter type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: Sample was treated with 0.001% glutaraldehyde for 20 min on ice prior quenching with 100 mM Tris-HCl pH 7.5 and gel filtration. |
|---|---|
| Source (natural) | Organism:  Influenza A virus (A/nt/60/1968(H3N2)) Influenza A virus (A/nt/60/1968(H3N2)) |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 250 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.35 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 / Component:
Details: Sample was purified in 20 mM HEPES, pH 7.5, 150 mM NaCl with Tween 20 added to a final concentration 0f 0.05% prior to plunging grids. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 3.5 sec before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 1178 / Average exposure time: 5.0 sec. / Average electron dose: 1.32 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 0.7 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 80 |
|---|
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)