[English] 日本語
 Yorodumi
Yorodumi- EMDB-40120: CryoEM structure of beta-2-adrenergic receptor in complex with Gs... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of beta-2-adrenergic receptor in complex with Gs heterotrimer, 10 sec after GTP addition (#5 of 20) | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / Adrenergic / Receptor / G protein / SIGNALING PROTEIN | |||||||||
| Function / homology | Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Isoform Gnas-2 of Guanine nucleotide-binding protein G(s) subunit alpha isoforms short Function and homology information Function and homology information | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Papasergi-Scott MM / Skiniotis G | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Time-resolved cryo-EM of G-protein activation by a GPCR. Authors: Makaía M Papasergi-Scott / Guillermo Pérez-Hernández / Hossein Batebi / Yang Gao / Gözde Eskici / Alpay B Seven / Ouliana Panova / Daniel Hilger / Marina Casiraghi / Feng He / Luis Maul ...Authors: Makaía M Papasergi-Scott / Guillermo Pérez-Hernández / Hossein Batebi / Yang Gao / Gözde Eskici / Alpay B Seven / Ouliana Panova / Daniel Hilger / Marina Casiraghi / Feng He / Luis Maul / Peter Gmeiner / Brian K Kobilka / Peter W Hildebrand / Georgios Skiniotis /    Abstract: G-protein-coupled receptors (GPCRs) activate heterotrimeric G proteins by stimulating guanine nucleotide exchange in the Gα subunit. To visualize this mechanism, we developed a time-resolved cryo-EM ...G-protein-coupled receptors (GPCRs) activate heterotrimeric G proteins by stimulating guanine nucleotide exchange in the Gα subunit. To visualize this mechanism, we developed a time-resolved cryo-EM approach that examines the progression of ensembles of pre-steady-state intermediates of a GPCR-G-protein complex. By monitoring the transitions of the stimulatory G protein in complex with the β-adrenergic receptor at short sequential time points after GTP addition, we identified the conformational trajectory underlying G-protein activation and functional dissociation from the receptor. Twenty structures generated from sequential overlapping particle subsets along this trajectory, compared to control structures, provide a high-resolution description of the order of main events driving G-protein activation in response to GTP binding. Structural changes propagate from the nucleotide-binding pocket and extend through the GTPase domain, enacting alterations to Gα switch regions and the α5 helix that weaken the G-protein-receptor interface. Molecular dynamics simulations with late structures in the cryo-EM trajectory support that enhanced ordering of GTP on closure of the α-helical domain against the nucleotide-bound Ras-homology domain correlates with α5 helix destabilization and eventual dissociation of the G protein from the GPCR. These findings also highlight the potential of time-resolved cryo-EM as a tool for mechanistic dissection of GPCR signalling events. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40120.map.gz emd_40120.map.gz | 230.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40120-v30.xml emd-40120-v30.xml emd-40120.xml emd-40120.xml | 21.3 KB 21.3 KB | Display Display |  EMDB header EMDB header |
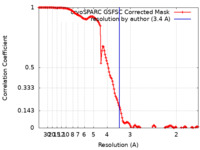
| FSC (resolution estimation) |  emd_40120_fsc.xml emd_40120_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_40120.png emd_40120.png | 94.9 KB | ||
| Masks |  emd_40120_msk_1.map emd_40120_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-40120.cif.gz emd-40120.cif.gz | 6.5 KB | ||
| Others |  emd_40120_additional_1.map.gz emd_40120_additional_1.map.gz emd_40120_half_map_1.map.gz emd_40120_half_map_1.map.gz emd_40120_half_map_2.map.gz emd_40120_half_map_2.map.gz | 122.2 MB 226.8 MB 226.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40120 http://ftp.pdbj.org/pub/emdb/structures/EMD-40120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40120 | HTTPS FTP |
-Related structure data
| Related structure data |  8gdzC  8ge1C  8ge2C  8ge3C  8ge4C  8ge5C  8ge6C  8ge7C  8ge8C  8ge9C  8geaC  8gebC  8gecC  8gedC  8geeC  8gefC  8gegC  8gehC  8geiC  8gejC  8gfvC  8gfwC  8gfxC  8gfyC  8gfzC  8gg0C  8gg1C  8gg2C  8gg3C  8gg4C  8gg5C  8gg6C  8gg7C  8gg8C  8gg9C  8ggaC  8ggbC  8ggcC  8ggeC  8ggfC  8ggiC  8ggjC  8ggkC  8gglC  8ggmC  8ggnC  8ggoC  8ggpC  8ggqC  8ggrC  8ggsC  8ggtC  8gguC  8ggvC  8ggwC  8ggxC  8ggyC  8ggzC  8gh0C  8gh1C  8unlC  8unmC  8unnC  8unoC  8unpC  8unqC  8unrC  8unsC  8untC  8unuC  8unvC  8unwC  8unxC  8unyC  8unzC  8uo0C  8uo1C  8uo2C  8uo3C  8uo4C C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40120.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40120.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8521 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40120_msk_1.map emd_40120_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened map
| File | emd_40120_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map
| File | emd_40120_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map
| File | emd_40120_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of beta-2 adrenergic receptor and Gs heterotrimer, 10 sec...
| Entire | Name: Complex of beta-2 adrenergic receptor and Gs heterotrimer, 10 sec post GTP treatment |
|---|---|
| Components |
|
-Supramolecule #1: Complex of beta-2 adrenergic receptor and Gs heterotrimer, 10 sec...
| Supramolecule | Name: Complex of beta-2 adrenergic receptor and Gs heterotrimer, 10 sec post GTP treatment type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKSTIVKQM RILHVNGFNG DSEKATKVQD IKNNLKEAIE TIVAAMSNLV PPVELANPEN QFRVDYILSV MNVPDFDFPP EFYEHAKALW EDEGVRACYE RSNEYQLIDC AQYFLDKIDV ...String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKSTIVKQM RILHVNGFNG DSEKATKVQD IKNNLKEAIE TIVAAMSNLV PPVELANPEN QFRVDYILSV MNVPDFDFPP EFYEHAKALW EDEGVRACYE RSNEYQLIDC AQYFLDKIDV IKQADYVPSD QDLLRCRVLT SGIFETKFQV DKVNFHMFDV GGQRDERRKW IQCFNDVTAI IFVVASSSYN MVIREDNQTN RLQEALNLFK SIWNNRWLRT ISVILFLNKQ DLLAEKVLAG KSKIEDYFPE FARYTTPEDA TPEPGEDPRV TRAKYFIRDE FLRISTASGD GRHYCYPHFT CAVDTENIRR VFNDCRDIIQ RMHLRQYELL UniProtKB: Isoform Gnas-2 of Guanine nucleotide-binding protein G(s) subunit alpha isoforms short |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GSSGSELDQL RQEAEQLKNQ IRDARKACAD ATLSQITNNI DPVGRIQMRT RRTLRGHLAK IYAMHWGTDS RLLVSASQDG KLIIWDSYTT NKVHAIPLRS SWVMTCAYAP SGNYVACGGL DNICSIYNLK TREGNVRVSR ELAGHTGYLS CCRFLDDNQI VTSSGDTTCA ...String: GSSGSELDQL RQEAEQLKNQ IRDARKACAD ATLSQITNNI DPVGRIQMRT RRTLRGHLAK IYAMHWGTDS RLLVSASQDG KLIIWDSYTT NKVHAIPLRS SWVMTCAYAP SGNYVACGGL DNICSIYNLK TREGNVRVSR ELAGHTGYLS CCRFLDDNQI VTSSGDTTCA LWDIETGQQT TTFTGHTGDV MSLSLAPDTR LFVSGACDAS AKLWDVREGM CRQTFTGHES DINAICFFPN GNAFATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGVTDDGMAV ATGSWDSFLK IWN |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L |
-Macromolecule #4: Beta-2 adrenergic receptor
| Macromolecule | Name: Beta-2 adrenergic receptor / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDAMGQPGN GSAFLLAPNR SHAPDHDVEN LYFQGTQQRD EVWVVGMGIV MSLIVLAIVF GNVLVITAIA KFERLQTVTN YFITSLACAD LVMGLAVVPF GAAHILTKTW TFGNFWCEFW TSIDVLCVTA SIETLCVIAV DRYFAITSPF ...String: MKTIIALSYI FCLVFADYKD DDDAMGQPGN GSAFLLAPNR SHAPDHDVEN LYFQGTQQRD EVWVVGMGIV MSLIVLAIVF GNVLVITAIA KFERLQTVTN YFITSLACAD LVMGLAVVPF GAAHILTKTW TFGNFWCEFW TSIDVLCVTA SIETLCVIAV DRYFAITSPF KYQSLLTKNK ARVIILMVWI VSGLTSFLPI QMHWYRATHQ EAINCYAEET CCDFFTNQAY AIASSIVSFY VPLVIMVFVY SRVFQEAKRQ LQKIDKSEGR FHVQNLSQVE QDGRTGHGLR RSSKFCLKEH KALKTLGIIM GTFTLCWLPF FIVNIVHVIQ DNLIRKEVYI LLNWIGYVNS GFNPLIYCRS PDFRIAFQEL LCLRRSSLKA YGNGYSSNGN TGEQSGLEVL FQGPYHVEQE KENKLLAEDL PGTEDFVGHQ GTVPSDNIDS QGRNASTNDS LLETSQVAPA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: UltrAuFoil R1.2/1.3 / Support film - Material: GOLD / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Average electron dose: 25.83 e/Å2 Details: Cryo-EM imaging of beta-2AR-Gs + GTP (10 sec) complex was performed on four separate grids over three collection sessions. The microscope was operated at 300 kV accelerating voltage, with a ...Details: Cryo-EM imaging of beta-2AR-Gs + GTP (10 sec) complex was performed on four separate grids over three collection sessions. The microscope was operated at 300 kV accelerating voltage, with a magnification at the camera of 58,679x in counting mode resulting in a magnified pixel size of 0.8521 Angstrom. For the first and second grids, movies were obtained at an exposure rate of 21.13 electrons/Angstrum^2/sec with defocus ranging from -0.4 - -2.0 micrometers. The total exposure time was 2.717 sec over 77 frames per movie stack. For an additional collection of the first grid, movies were obtained at an exposure rate of 20.95 electrons/ Angstrum^2/sec with defocus ranging from -0.4 -2.0 micrometers. The total exposure time was 2.717 sec over 77 frames per movie stack. For the third and fourth grids, movies were obtained at an exposure rate of 30.71 electrons/ Angstrum^2/sec with defocus ranging from -0.5 - -1.6 micrometers. The total exposure time was 2.008 sec over 79 frames per movie stack. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


































































































































































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)