[English] 日本語
 Yorodumi
Yorodumi- EMDB-3817: Structure of an RNA polymerase II-DSIF transcription elongation c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3817 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of an RNA polymerase II-DSIF transcription elongation complex, DSIF-EC3 | ||||||||||||
 Map data Map data | Locally filtered map, B-factor 0 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | RNA polymerase II / transcription elongation / TRANSCRIPTION | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationFormation of RNA Pol II elongation complex / Formation of the Early Elongation Complex / Transcriptional regulation by small RNAs / FGFR2 alternative splicing / RNA polymerase II transcribes snRNA genes / mRNA Capping / mRNA Splicing - Minor Pathway / RNA Polymerase I Transcription Initiation / RNA Polymerase I Promoter Escape / RNA Polymerase II Promoter Escape ...Formation of RNA Pol II elongation complex / Formation of the Early Elongation Complex / Transcriptional regulation by small RNAs / FGFR2 alternative splicing / RNA polymerase II transcribes snRNA genes / mRNA Capping / mRNA Splicing - Minor Pathway / RNA Polymerase I Transcription Initiation / RNA Polymerase I Promoter Escape / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase I Transcription Termination / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA Polymerase III Transcription Initiation From Type 1 Promoter / RNA Polymerase III Transcription Initiation From Type 2 Promoter / RNA Polymerase III Transcription Initiation From Type 3 Promoter / RNA Pol II CTD phosphorylation and interaction with CE / Estrogen-dependent gene expression / RNA Polymerase II Pre-transcription Events / TP53 Regulates Transcription of DNA Repair Genes / RNA Polymerase II Transcription Elongation / Processing of Capped Intron-Containing Pre-mRNA / B-WICH complex positively regulates rRNA expression / mRNA Splicing - Major Pathway / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / negative regulation of DNA-templated transcription, elongation / DNA/RNA hybrid binding / DSIF complex / regulation of transcription elongation by RNA polymerase II / transcription elongation-coupled chromatin remodeling / positive regulation of DNA-templated transcription, elongation / Abortive elongation of HIV-1 transcript in the absence of Tat / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / RNA polymerase II complex binding / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / negative regulation of transcription elongation by RNA polymerase II / RNA polymerase II transcribes snRNA genes / positive regulation of macroautophagy / Pausing and recovery of Tat-mediated HIV elongation / Tat-mediated HIV elongation arrest and recovery / positive regulation of translational initiation / HIV elongation arrest and recovery / Pausing and recovery of HIV elongation / nuclear-transcribed mRNA catabolic process / Tat-mediated elongation of the HIV-1 transcript / core promoter sequence-specific DNA binding / RNA polymerase I complex / Formation of HIV-1 elongation complex containing HIV-1 Tat / RNA polymerase III complex / transcription elongation by RNA polymerase I / Formation of HIV elongation complex in the absence of HIV Tat / RNA polymerase II, core complex / tRNA transcription by RNA polymerase III / transcription by RNA polymerase I / RNA Polymerase II Transcription Elongation / Formation of RNA Pol II elongation complex / translation initiation factor binding / transcription-coupled nucleotide-excision repair / RNA Polymerase II Pre-transcription Events / TP53 Regulates Transcription of DNA Repair Genes / transcription initiation at RNA polymerase II promoter / transcription elongation by RNA polymerase II / positive regulation of transcription elongation by RNA polymerase II / P-body / euchromatin / protein-DNA complex / mRNA transcription by RNA polymerase II / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / single-stranded DNA binding / Hydrolases; Acting on ester bonds; Exoribonucleases producing 5'-phosphomonoesters / transcription by RNA polymerase II / nucleic acid binding / chromosome, telomeric region / protein dimerization activity / single-stranded RNA binding / protein heterodimerization activity / RNA-directed RNA polymerase / nucleotide binding / hydrolase activity / RNA-directed RNA polymerase activity / mRNA binding / chromatin binding / regulation of DNA-templated transcription / nucleolus / magnesium ion binding / enzyme binding / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / DNA binding / RNA binding Similarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | ||||||||||||
 Authors Authors | Bernecky C / Plitzko JM / Cramer P | ||||||||||||
| Funding support |  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2017 Journal: Nat Struct Mol Biol / Year: 2017Title: Structure of a transcribing RNA polymerase II-DSIF complex reveals a multidentate DNA-RNA clamp. Authors: Carrie Bernecky / Jürgen M Plitzko / Patrick Cramer /  Abstract: During transcription, RNA polymerase II (Pol II) associates with the conserved elongation factor DSIF. DSIF renders the elongation complex stable and functions during Pol II pausing and RNA ...During transcription, RNA polymerase II (Pol II) associates with the conserved elongation factor DSIF. DSIF renders the elongation complex stable and functions during Pol II pausing and RNA processing. We combined cryo-EM and X-ray crystallography to determine the structure of the mammalian Pol II-DSIF elongation complex at a nominal resolution of 3.4 Å. Human DSIF has a modular structure with two domains forming a DNA clamp, two domains forming an RNA clamp, and one domain buttressing the RNA clamp. The clamps maintain the transcription bubble, position upstream DNA, and retain the RNA transcript in the exit tunnel. The mobile C-terminal region of DSIF is located near exiting RNA, where it can recruit factors for RNA processing. The structure provides insight into the roles of DSIF during mRNA synthesis. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3817.map.gz emd_3817.map.gz | 58.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3817-v30.xml emd-3817-v30.xml emd-3817.xml emd-3817.xml | 47.5 KB 47.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3817.png emd_3817.png | 106.2 KB | ||
| Masks |  emd_3817_msk_1.map emd_3817_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-3817.cif.gz emd-3817.cif.gz | 11.4 KB | ||
| Others |  emd_3817_additional_1.map.gz emd_3817_additional_1.map.gz emd_3817_additional_2.map.gz emd_3817_additional_2.map.gz emd_3817_half_map_1.map.gz emd_3817_half_map_1.map.gz emd_3817_half_map_2.map.gz emd_3817_half_map_2.map.gz | 60 MB 59.6 MB 49.6 MB 49.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3817 http://ftp.pdbj.org/pub/emdb/structures/EMD-3817 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3817 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3817 | HTTPS FTP |
-Related structure data
| Related structure data |  5oikMC  3815C  3816C  3818C  3819C  5ohoC  5ohqC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3817.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3817.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
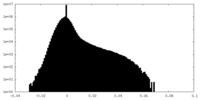
| Annotation | Locally filtered map, B-factor 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
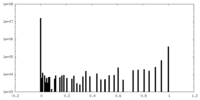
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_3817_msk_1.map emd_3817_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
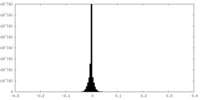
| Density Histograms |
-Additional map: None
| File | emd_3817_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
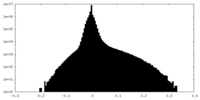
| Density Histograms |
-Additional map: None
| File | emd_3817_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
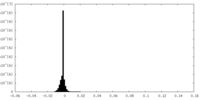
| Density Histograms |
-Half map: Halfmap 1
| File | emd_3817_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap 2
| File | emd_3817_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : RNA polymerase II-DSIF elongation complex
+Supramolecule #1: RNA polymerase II-DSIF elongation complex
+Supramolecule #2: RNA polymerase II
+Supramolecule #3: DSIF
+Supramolecule #4: DNA-RNA synthetic construct
+Macromolecule #1: DNA-directed RNA polymerase II subunit RPB1
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: DNA-directed RNA polymerase II subunit RPB3
+Macromolecule #4: Polymerase (RNA) II (DNA directed) polypeptide D
+Macromolecule #5: DNA-directed RNA polymerases I, II, and III subunit RPABC1
+Macromolecule #6: DNA-directed RNA polymerases I, II, and III subunit RPABC2
+Macromolecule #7: DNA-directed RNA polymerase II subunit RPB7
+Macromolecule #8: Uncharacterized protein
+Macromolecule #9: DNA-directed RNA polymerase II subunit RPB9
+Macromolecule #10: DNA-directed RNA polymerases I, II, and III subunit RPABC5
+Macromolecule #11: DNA-directed RNA polymerase II subunit RPB11
+Macromolecule #12: DNA-directed RNA polymerases I, II, and III subunit RPABC4
+Macromolecule #16: Transcription elongation factor SPT4
+Macromolecule #17: Transcription elongation factor SPT5
+Macromolecule #13: DNA (43-MER)
+Macromolecule #15: DNA (43-MER)
+Macromolecule #14: RNA (5'-R(P*UP*AP*UP*AP*UP*AP*CP*AP*UP*AP*AP*AP*GP*AP*CP*CP*AP*GP...
+Macromolecule #18: ZINC ION
+Macromolecule #19: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.25 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.25 Component:
Details: pH was adjusted at 25 degrees Celsius | |||||||||||||||
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: Sample (four microliters) was applied to glow-discharged Quantifoil R 2/1 holey carbon grids, which were then blotted for 8.5s and plunge-frozen in liquid ethane.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum / Energy filter - Lower energy threshold: 0 eV / Energy filter - Upper energy threshold: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-33 / Number grids imaged: 1 / Number real images: 2549 / Average exposure time: 9.9 sec. / Average electron dose: 33.0 e/Å2 Details: Movies were aligned and binned to the physical pixel size of 1.35 angstroms. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 3.6 µm / Calibrated defocus min: 0.6 µm / Calibrated magnification: 37037 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.6 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 37000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-5oik: |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)