[English] 日本語
 Yorodumi
Yorodumi- EMDB-37976: Human FL Metabotropic glutamate receptor 5, mGlu5-5M with agonist... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human FL Metabotropic glutamate receptor 5, mGlu5-5M with agonist and PAM, W785A mutant | ||||||||||||
 Map data Map data | Combined sharpened map with B-factor of -62 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | G-PROTEIN COUPLED RECEPTORS / SIGNAL TRANSDUCTION / METABOTROPIC GLUTAMATE RECEPTOR / Agonist / active state / MEMBRANE PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationA2A adenosine receptor binding / sensory perception of hot stimulus / negative regulation of dendritic spine morphogenesis / : / G protein-coupled receptor activity involved in regulation of postsynaptic membrane potential / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / operant conditioning / protein localization to nuclear inner membrane / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / positive regulation of cellular response to hypoxia ...A2A adenosine receptor binding / sensory perception of hot stimulus / negative regulation of dendritic spine morphogenesis / : / G protein-coupled receptor activity involved in regulation of postsynaptic membrane potential / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / operant conditioning / protein localization to nuclear inner membrane / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / positive regulation of cellular response to hypoxia / positive regulation of long-term neuronal synaptic plasticity / positive regulation of sensory perception of pain / negative regulation of excitatory postsynaptic potential / positive regulation of dopamine secretion / desensitization of G protein-coupled receptor signaling pathway / G protein-coupled glutamate receptor signaling pathway / positive regulation of neural precursor cell proliferation / Class C/3 (Metabotropic glutamate/pheromone receptors) / nuclear inner membrane / glutamate receptor activity / Neurexins and neuroligins / astrocyte projection / response to corticosterone / response to morphine / temperature homeostasis / protein tyrosine kinase activator activity / conditioned place preference / regulation of synaptic transmission, glutamatergic / regulation of long-term synaptic depression / positive regulation of calcium-mediated signaling / protein tyrosine kinase binding / dendritic shaft / response to amphetamine / locomotory behavior / synapse organization / postsynaptic density membrane / G protein-coupled receptor activity / Schaffer collateral - CA1 synapse / cognition / cellular response to amyloid-beta / positive regulation of cytosolic calcium ion concentration / dendritic spine / G alpha (q) signalling events / chemical synaptic transmission / response to ethanol / learning or memory / positive regulation of MAPK cascade / response to antibiotic / neuronal cell body / regulation of DNA-templated transcription / dendrite / glutamatergic synapse / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | ||||||||||||
 Authors Authors | Vinothkumar KR / Lebon G / Cannone G | ||||||||||||
| Funding support |  India, India,  France, 3 items France, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2025 Journal: Nat Commun / Year: 2025Title: Conformational diversity in class C GPCR positive allosteric modulation. Authors: Giuseppe Cannone / Ludovic Berto / Fanny Malhaire / Gavin Ferguson / Aurelien Fouillen / Stéphanie Balor / Joan Font-Ingles / Amadeu Llebaria / Cyril Goudet / Abhay Kotecha / Vinothkumar K ...Authors: Giuseppe Cannone / Ludovic Berto / Fanny Malhaire / Gavin Ferguson / Aurelien Fouillen / Stéphanie Balor / Joan Font-Ingles / Amadeu Llebaria / Cyril Goudet / Abhay Kotecha / Vinothkumar K R / Guillaume Lebon /      Abstract: The metabotropic glutamate receptors (mGlus) are class C G protein-coupled receptors (GPCR) that form obligate dimers activated by the major excitatory neurotransmitter L-glutamate. The architecture ...The metabotropic glutamate receptors (mGlus) are class C G protein-coupled receptors (GPCR) that form obligate dimers activated by the major excitatory neurotransmitter L-glutamate. The architecture of mGlu receptor comprises an extracellular Venus-Fly Trap domain (VFT) connected to the transmembrane domain (7TM) through a Cysteine-Rich Domain (CRD). The binding of L-glutamate in the VFTs and subsequent conformational change results in the signal being transmitted to the 7TM inducing G protein binding and activation. The mGlu receptors signal transduction can be allosterically potentiated by positive allosteric modulators (PAMs) binding to the 7TMs, which are of therapeutic interest in various neurological disorders. Here, we report the cryoEM structures of metabotropic glutamate receptor 5 (mGlu) purified with three chemically and pharmacologically distinct PAMs. We find that the PAMs modulate the receptor equilibrium through their different binding modes, revealing how their interactions in the 7TMs impact the mGlu receptor conformational landscape and function. In addition, we identified a PAM-free but agonist-bound intermediate state that also reveals interactions mediated by intracellular loop 2. The activation of mGlu receptor is a multi-step process in which the binding of the PAMs in the 7TM modulates the equilibrium towards the active state. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37976.map.gz emd_37976.map.gz | 200.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37976-v30.xml emd-37976-v30.xml emd-37976.xml emd-37976.xml | 23.4 KB 23.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37976_fsc.xml emd_37976_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_37976.png emd_37976.png | 108.3 KB | ||
| Filedesc metadata |  emd-37976.cif.gz emd-37976.cif.gz | 7.8 KB | ||
| Others |  emd_37976_half_map_1.map.gz emd_37976_half_map_1.map.gz emd_37976_half_map_2.map.gz emd_37976_half_map_2.map.gz | 200.6 MB 200.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37976 http://ftp.pdbj.org/pub/emdb/structures/EMD-37976 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37976 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37976 | HTTPS FTP |
-Related structure data
| Related structure data |  8x0eMC  8x0bC  8x0cC  8x0dC  8x0fC  8x0gC  8x0hC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_37976.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37976.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Combined sharpened map with B-factor of -62 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: One of the half map from Cryosparc
| File | emd_37976_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of the half map from Cryosparc | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: One of the half map from Cryosparc
| File | emd_37976_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of the half map from Cryosparc | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Thermostabilised full length human mGluR5, W785A mutant
| Entire | Name: Thermostabilised full length human mGluR5, W785A mutant |
|---|---|
| Components |
|
-Supramolecule #1: Thermostabilised full length human mGluR5, W785A mutant
| Supramolecule | Name: Thermostabilised full length human mGluR5, W785A mutant type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: Receptor was purified with orthosteric Quisqualate and PAM VU0424465 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 200 KDa |
-Macromolecule #1: Metabotropic glutamate receptor 5
| Macromolecule | Name: Metabotropic glutamate receptor 5 / type: protein_or_peptide / ID: 1 Details: The N-terminal sequence (DYKDDDDKHHHHHHHHHHLEVLFQGP) is the tag and linker), which has been cleaved by protease before structural experiment. Here, it is included for completion. The ...Details: The N-terminal sequence (DYKDDDDKHHHHHHHHHHLEVLFQGP) is the tag and linker), which has been cleaved by protease before structural experiment. Here, it is included for completion. The sequence when compared to uniprot starts at 21 and ends at 856 (construct used for expression). The construct has the following mutations (with reference to Uniprot ID P41594) H350L - mutation for nanobody binding T742A, S753A, T777A, I799A, A813L (thermostabilising mutant) W785A - introduced mutant to test for PAM binding. There are 10 disulfide bonds per chain and 2 NAG molecules. Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 93.619383 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSSERRVVAH MPGDIIIGAL FSVHHQPTVD KVHERKCGAV REQYGIQRVE AMLHTLERIN SDPTLLPNIT LGCEIRDSCW HSAVALEQS IEFIRDSLIS SEEEEGLVRC VDGSSSSFRS KKPIVGVIGP GSSSVAIQVQ NLLQLFNIPQ IAYSATSMDL S DKTLFKYF ...String: QSSERRVVAH MPGDIIIGAL FSVHHQPTVD KVHERKCGAV REQYGIQRVE AMLHTLERIN SDPTLLPNIT LGCEIRDSCW HSAVALEQS IEFIRDSLIS SEEEEGLVRC VDGSSSSFRS KKPIVGVIGP GSSSVAIQVQ NLLQLFNIPQ IAYSATSMDL S DKTLFKYF MRVVPSDAQQ ARAMVDIVKR YNWTYVSAVH TEGNYGESGM EAFKDMSAKE GICIAHSYKI YSNAGEQSFD KL LKKLTSH LPKARVVACF CEGMTVRGLL MAMRRLGLAG EFLLLGSDGW ADRYDVTDGY QREAVGGITI KLQSPDVKWF DDY YLKLRP ETNLRNPWFQ EFWQHRFQCR LEGFPQENSK YNKTCNSSLT LKTHHVQDSK MGFVINAIYS MAYGLHNMQM SLCP GYAGL CDAMKPIDGR KLLESLMKTN FTGVSGDTIL FDENGDSPGR YEIMNFKEMG KDYFDYINVG SWDNGELKMD DDEVW SKKS NIIRSVCSEP CEKGQIKVIR KGEVSCCWTC TPCKENEYVF DEYTCKACQL GSWPTDDLTG CDLIPVQYLR WGDPEP IAA VVFACLGLLA TLFVTVVFII YRDTPVVKSS SRELCYIILA GICLGYLCTF CLIAKPKQIY CYLQRIGIGL SPAMSYS AL VTKTNRIARI LAGSKKKICT KKPRFMSACA QLVIAFILIC IQLGIIVALF IMEPPDIMHD YPSIREVYLI CNTTNLGV V APLGYNGLLI LACTFYAFKT RNVPANFNEA KYIAFAMYTT CIIALAFVPI YFGSNYKAIT MCFSVSLSAT VLLGCMFVP KVYIILAKPE RNVRSAFTTS TVVRMHVGDG KSSSAA UniProtKB: Metabotropic glutamate receptor 5 |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #3: (S)-2-AMINO-3-(3,5-DIOXO-[1,2,4]OXADIAZOLIDIN-2-YL)-PROPIONIC ACID
| Macromolecule | Name: (S)-2-AMINO-3-(3,5-DIOXO-[1,2,4]OXADIAZOLIDIN-2-YL)-PROPIONIC ACID type: ligand / ID: 3 / Number of copies: 2 / Formula: QUS |
|---|---|
| Molecular weight | Theoretical: 189.126 Da |
| Chemical component information | 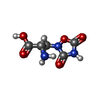 ChemComp-QUS: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 289 K / Instrument: FEI VITROBOT MARK IV / Details: Blot force was set to 0. | |||||||||||||||
| Details | Receptor is purified in detergent micelles and monodisperse. The receptor is purified with 10 uM Quisqualate and 10 uM PAM VU0424465 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 2 / Number real images: 5762 / Average exposure time: 60.0 sec. / Average electron dose: 28.25 e/Å2 Details: Images were collected in movie mode for 60 seconds and 25 frames were stored. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 130841 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.8 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Source name: Other / Chain - Initial model type: experimental model Details: The initial model was from this deposition ID, which has been submitted |
|---|---|
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Overall B value: 183.8 |
| Output model |  PDB-8x0e: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)






































