[English] 日本語
 Yorodumi
Yorodumi- EMDB-31537: Thermostabilised full length human mGluR5-5M with orthosteric ant... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31537 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Thermostabilised full length human mGluR5-5M with orthosteric antagonist, LY341495 | ||||||||||||||||||
 Map data Map data | Post processed map from Relion, with a B-factor sharpening of -82 | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | G-protein coupled receptors / signal transduction / metabotropic glutamate receptor / inactive state / MEMBRANE PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationA2A adenosine receptor binding / sensory perception of hot stimulus / : / negative regulation of dendritic spine morphogenesis / G protein-coupled receptor activity involved in regulation of postsynaptic membrane potential / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / operant conditioning / protein localization to nuclear inner membrane / positive regulation of cellular response to hypoxia ...A2A adenosine receptor binding / sensory perception of hot stimulus / : / negative regulation of dendritic spine morphogenesis / G protein-coupled receptor activity involved in regulation of postsynaptic membrane potential / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / operant conditioning / protein localization to nuclear inner membrane / positive regulation of cellular response to hypoxia / positive regulation of long-term neuronal synaptic plasticity / positive regulation of sensory perception of pain / negative regulation of excitatory postsynaptic potential / positive regulation of dopamine secretion / desensitization of G protein-coupled receptor signaling pathway / G protein-coupled glutamate receptor signaling pathway / positive regulation of neural precursor cell proliferation / Class C/3 (Metabotropic glutamate/pheromone receptors) / glutamate receptor activity / nuclear inner membrane / Neurexins and neuroligins / astrocyte projection / response to corticosterone / response to morphine / temperature homeostasis / protein tyrosine kinase activator activity / conditioned place preference / regulation of synaptic transmission, glutamatergic / regulation of long-term synaptic depression / positive regulation of calcium-mediated signaling / protein tyrosine kinase binding / dendritic shaft / response to amphetamine / locomotory behavior / synapse organization / postsynaptic density membrane / G protein-coupled receptor activity / Schaffer collateral - CA1 synapse / cognition / cellular response to amyloid-beta / positive regulation of cytosolic calcium ion concentration / G alpha (q) signalling events / dendritic spine / chemical synaptic transmission / response to ethanol / learning or memory / positive regulation of MAPK cascade / response to antibiotic / neuronal cell body / regulation of DNA-templated transcription / dendrite / glutamatergic synapse / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | ||||||||||||||||||
 Authors Authors | Vinothkumar KR / Cannone G | ||||||||||||||||||
| Funding support |  India, India,  United Kingdom, United Kingdom,  France, 5 items France, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2021 Journal: Cell Rep / Year: 2021Title: Agonists and allosteric modulators promote signaling from different metabotropic glutamate receptor 5 conformations. Authors: Chady Nasrallah / Giuseppe Cannone / Julie Briot / Karine Rottier / Alice E Berizzi / Chia-Ying Huang / Robert B Quast / Francois Hoh / Jean-Louis Banères / Fanny Malhaire / Ludovic Berto / ...Authors: Chady Nasrallah / Giuseppe Cannone / Julie Briot / Karine Rottier / Alice E Berizzi / Chia-Ying Huang / Robert B Quast / Francois Hoh / Jean-Louis Banères / Fanny Malhaire / Ludovic Berto / Anaëlle Dumazer / Joan Font-Ingles / Xavier Gómez-Santacana / Juanlo Catena / Julie Kniazeff / Cyril Goudet / Amadeu Llebaria / Jean-Philippe Pin / Kutti R Vinothkumar / Guillaume Lebon /      Abstract: Metabotropic glutamate receptors (mGluRs) are dimeric G-protein-coupled receptors activated by the main excitatory neurotransmitter, L-glutamate. mGluR activation by agonists binding in the venus ...Metabotropic glutamate receptors (mGluRs) are dimeric G-protein-coupled receptors activated by the main excitatory neurotransmitter, L-glutamate. mGluR activation by agonists binding in the venus flytrap domain is regulated by positive (PAM) or negative (NAM) allosteric modulators binding to the 7-transmembrane domain (7TM). We report the cryo-electron microscopy structures of fully inactive and intermediate-active conformations of mGlu receptor bound to an antagonist and a NAM or an agonist and a PAM, respectively, as well as the crystal structure of the 7TM bound to a photoswitchable NAM. The agonist induces a large movement between the subunits, bringing the 7TMs together and stabilizing a 7TM conformation structurally similar to the inactive state. Using functional approaches, we demonstrate that the PAM stabilizes a 7TM active conformation independent of the conformational changes induced by agonists, representing an alternative mode of mGlu activation. These findings provide a structural basis for different mGluR activation modes. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31537.map.gz emd_31537.map.gz | 93.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31537-v30.xml emd-31537-v30.xml emd-31537.xml emd-31537.xml | 27.3 KB 27.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_31537_fsc.xml emd_31537_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_31537.png emd_31537.png | 44.1 KB | ||
| Filedesc metadata |  emd-31537.cif.gz emd-31537.cif.gz | 7.8 KB | ||
| Others |  emd_31537_additional_1.map.gz emd_31537_additional_1.map.gz emd_31537_additional_2.map.gz emd_31537_additional_2.map.gz emd_31537_half_map_1.map.gz emd_31537_half_map_1.map.gz emd_31537_half_map_2.map.gz emd_31537_half_map_2.map.gz | 92 MB 95 MB 79.3 MB 79.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31537 http://ftp.pdbj.org/pub/emdb/structures/EMD-31537 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31537 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31537 | HTTPS FTP |
-Related structure data
| Related structure data |  7fd9MC  7fd8C  7p2lC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31537.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31537.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post processed map from Relion, with a B-factor sharpening of -82 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.89 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Postprocessed map sharpened with a B factor of...
| File | emd_31537_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed map sharpened with a B factor of -20, used in model building | ||||||||||||
| Projections & Slices |
| ||||||||||||
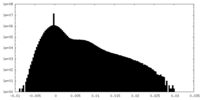
| Density Histograms |
-Additional map: Postprocessed map sharpened with a B factor of...
| File | emd_31537_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed map sharpened with a B factor of -164, used in model building | ||||||||||||
| Projections & Slices |
| ||||||||||||
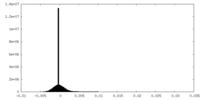
| Density Histograms |
-Half map: One of the half map from refinement
| File | emd_31537_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of the half map from refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
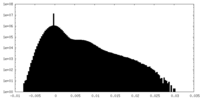
| Density Histograms |
-Half map: One of the half map from refinement
| File | emd_31537_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of the half map from refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human FL mGluR5-5M with orthosteric antagonist LY341495 and NAM MPEP
| Entire | Name: Human FL mGluR5-5M with orthosteric antagonist LY341495 and NAM MPEP |
|---|---|
| Components |
|
-Supramolecule #1: Human FL mGluR5-5M with orthosteric antagonist LY341495 and NAM MPEP
| Supramolecule | Name: Human FL mGluR5-5M with orthosteric antagonist LY341495 and NAM MPEP type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 194 KDa |
-Macromolecule #1: Metabotropic glutamate receptor 5
| Macromolecule | Name: Metabotropic glutamate receptor 5 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 96.953977 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DYKDDDDKHH HHHHHHHHLE VLFQGPQSSE RRVVAHMPGD IIIGALFSVH HQPTVDKVHE RKCGAVREQY GIQRVEAMLH TLERINSDP TLLPNITLGC EIRDSCWHSA VALEQSIEFI RDSLISSEEE EGLVRCVDGS SSSFRSKKPI VGVIGPGSSS V AIQVQNLL ...String: DYKDDDDKHH HHHHHHHHLE VLFQGPQSSE RRVVAHMPGD IIIGALFSVH HQPTVDKVHE RKCGAVREQY GIQRVEAMLH TLERINSDP TLLPNITLGC EIRDSCWHSA VALEQSIEFI RDSLISSEEE EGLVRCVDGS SSSFRSKKPI VGVIGPGSSS V AIQVQNLL QLFNIPQIAY SATSMDLSDK TLFKYFMRVV PSDAQQARAM VDIVKRYNWT YVSAVHTEGN YGESGMEAFK DM SAKEGIC IAHSYKIYSN AGEQSFDKLL KKLTSHLPKA RVVACFCEGM TVRGLLMAMR RLGLAGEFLL LGSDGWADRY DVT DGYQRE AVGGITIKLQ SPDVKWFDDY YLKLRPETNL RNPWFQEFWQ HRFQCRLEGF PQENSKYNKT CNSSLTLKTH HVQD SKMGF VINAIYSMAY GLHNMQMSLC PGYAGLCDAM KPIDGRKLLE SLMKTAFTGV SGDTILFDEN GDSPGRYEIM NFKEM GKDY FDYINVGSWD NGELKMDDDE VWSKKSNIIR SVCSEPCEKG QIKVIRKGEV SCCWTCTPCK ENEYVFDEYT CKACQL GSW PTDDLTGCDL IPVQYLRWGD PEPIAAVVFA CLGLLATLFV TVVFIIYRDT PVVKSSSREL CYIILAGICL GYLCTFC LI AKPKQIYCYL QRIGIGLSPA MSYSALVTKT NRIARILAGS KKKICTKKPR FMSACAQLVI AFILICIQLG IIVALFIM E PPDIMHDYPS IREVYLICNT TNLGVVAPLG YNGLLILACT FYAFKTRNVP ANFNEAKYIA FAMYTTCIIW LAFVPIYFG SNYKAITMCF SVSLSATVLL GCMFVPKVYI ILAKPERNVR SAFTTSTVVR MHVGDGKSSS AA UniProtKB: Metabotropic glutamate receptor 5 |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #3: 2-[(1S,2S)-2-carboxycyclopropyl]-3-(9H-xanthen-9-yl)-D-alanine
| Macromolecule | Name: 2-[(1S,2S)-2-carboxycyclopropyl]-3-(9H-xanthen-9-yl)-D-alanine type: ligand / ID: 3 / Number of copies: 2 / Formula: Z99 |
|---|---|
| Molecular weight | Theoretical: 353.369 Da |
| Chemical component information |  ChemComp-Z99: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||
| Grid | Model: Quantifoil R0.6/1 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.02 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV / Details: Blot force was 10, 3.5 seconds blotting time. | |||||||||
| Details | mGluR5 with orthosteric antagonist LY341495 and NAM (MPEP) purified in DDM/CHS |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV Details: Imaging was done in EFTEM mode. Images were collected using beam shift in a series of 3x3 holes before stage shift. Active beam tilt compensation was turned on during the acquisition. |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Number grids imaged: 1 / Number real images: 16668 / Average exposure time: 5.2 sec. / Average electron dose: 49.9 e/Å2 Details: Dose rate was 7.6 e/p/s. Total number of frames is 32 and each frame has a dose of 1.56 e/A2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 44500 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Details | Refinement was performed with jelly body restraints | ||||||
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Overall B value: 147.5 | ||||||
| Output model |  PDB-7fd9: |
 Movie
Movie Controller
Controller


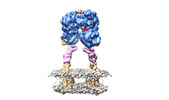








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)