[English] 日本語
 Yorodumi
Yorodumi- EMDB-31208: Local maps of TFIID-based PIC-Mediator complex (deletion of MED1-IDR) -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Local maps of TFIID-based PIC-Mediator complex (deletion of MED1-IDR) | |||||||||
 Map data Map data | CAK module in the TFIID-based PIC-Mediator complex (deletion of MED1-IDR) | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.54 Å | |||||||||
 Authors Authors | Chen X / Qi Y / Wang X / Wu Z / Yin X / Li J / Liu W / Xu Y | |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Structures of the human Mediator and Mediator-bound preinitiation complex. Authors: Xizi Chen / Xiaotong Yin / Jiabei Li / Zihan Wu / Yilun Qi / Xinxin Wang / Weida Liu / Yanhui Xu /  Abstract: The 1.3-megadalton transcription factor IID (TFIID) is required for preinitiation complex (PIC) assembly and RNA polymerase II (Pol II)-mediated transcription initiation on almost all genes. The 26- ...The 1.3-megadalton transcription factor IID (TFIID) is required for preinitiation complex (PIC) assembly and RNA polymerase II (Pol II)-mediated transcription initiation on almost all genes. The 26-subunit Mediator stimulates transcription and cyclin-dependent kinase 7 (CDK7)-mediated phosphorylation of the Pol II C-terminal domain (CTD). We determined the structures of human Mediator in the Tail module-extended (at near-atomic resolution) and Tail-bent conformations and structures of TFIID-based PIC-Mediator (76 polypeptides, ~4.1 megadaltons) in four distinct conformations. PIC-Mediator assembly induces concerted reorganization (Head-tilting and Middle-down) of Mediator and creates a Head-Middle sandwich, which stabilizes two CTD segments and brings CTD to CDK7 for phosphorylation; this suggests a CTD-gating mechanism favorable for phosphorylation. The TFIID-based PIC architecture modulates Mediator organization and TFIIH stabilization, underscoring the importance of TFIID in orchestrating PIC-Mediator assembly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31208.map.gz emd_31208.map.gz | 15.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31208-v30.xml emd-31208-v30.xml emd-31208.xml emd-31208.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31208.png emd_31208.png | 49.4 KB | ||
| Others |  emd_31208_additional_1.map.gz emd_31208_additional_1.map.gz emd_31208_additional_2.map.gz emd_31208_additional_2.map.gz emd_31208_additional_3.map.gz emd_31208_additional_3.map.gz | 227.9 MB 3.1 MB 1.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31208 http://ftp.pdbj.org/pub/emdb/structures/EMD-31208 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31208 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31208 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_31208.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31208.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CAK module in the TFIID-based PIC-Mediator complex (deletion of MED1-IDR) | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.405 Å | ||||||||||||||||||||||||||||||||||||
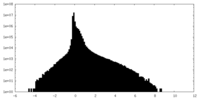
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Mediator module in the TFIID-based PIC-Mediator complex (deletion...
| File | emd_31208_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mediator module in the TFIID-based PIC-Mediator complex (deletion of MED1-IDR) | ||||||||||||
| Projections & Slices |
| ||||||||||||
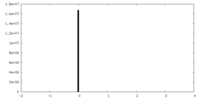
| Density Histograms |
-Additional map: Pol II module in the TFIID-based PIC-Mediator complex...
| File | emd_31208_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Pol II module in the TFIID-based PIC-Mediator complex (deletion of MED1-IDR) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: CTD-Mediator in the TFIID-based PIC-Mediator complex (deletion of...
| File | emd_31208_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CTD-Mediator in the TFIID-based PIC-Mediator complex (deletion of MED1-IDR) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Local maps of TFIID-based PIC-Mediator complex (deletion of MED1-IDR)
| Entire | Name: Local maps of TFIID-based PIC-Mediator complex (deletion of MED1-IDR) |
|---|---|
| Components |
|
-Supramolecule #1: Local maps of TFIID-based PIC-Mediator complex (deletion of MED1-IDR)
| Supramolecule | Name: Local maps of TFIID-based PIC-Mediator complex (deletion of MED1-IDR) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#72 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.54 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 110227 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)