+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30018 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the BAM complex | |||||||||
 Map data Map data | Structure of the BAM complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | b-barrel assembly machinery (BAM) complex / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationBam protein complex / Gram-negative-bacterium-type cell outer membrane assembly / protein insertion into membrane / Secretion of toxins / cell outer membrane / protein-folding chaperone binding / protein-macromolecule adaptor activity / cell adhesion / response to antibiotic / cell surface ...Bam protein complex / Gram-negative-bacterium-type cell outer membrane assembly / protein insertion into membrane / Secretion of toxins / cell outer membrane / protein-folding chaperone binding / protein-macromolecule adaptor activity / cell adhesion / response to antibiotic / cell surface / membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Xiao L / Huang Y | |||||||||
 Citation Citation |  Journal: FASEB J / Year: 2021 Journal: FASEB J / Year: 2021Title: Structures of the β-barrel assembly machine recognizing outer membrane protein substrates. Authors: Le Xiao / Long Han / Bufan Li / Manfeng Zhang / Haizhen Zhou / Qingshan Luo / Xinzheng Zhang / Yihua Huang /  Abstract: β-barrel outer membrane proteins (β-OMPs) play critical roles in nutrition acquisition, protein import/export, and other fundamental biological processes. The assembly of β-OMPs in Gram-negative ...β-barrel outer membrane proteins (β-OMPs) play critical roles in nutrition acquisition, protein import/export, and other fundamental biological processes. The assembly of β-OMPs in Gram-negative bacteria is mediated by the β-barrel assembly machinery (BAM) complex, yet its precise mechanism remains elusive. Here, we report two structures of the BAM complex in detergents and in nanodisks, and two crystal structures of the BAM complex with bound substrates. Structural analysis indicates that the membrane compositions surrounding the BAM complex could modulate its overall conformations, indicating low energy barriers between different conformational states and a highly dynamic nature of the BAM complex. Importantly, structures of the BAM complex with bound substrates and the related functional analysis show that the first β-strand of the BamA β-barrel (β1 ) in the BAM complex is associated with the last but not the first β-strand of a β-OMP substrate via antiparallel β-strand interactions. These observations are consistent with the β-signal hypothesis during β-OMP biogenesis, and suggest that the β1 strand in the BAM complex may interact with the last β-strand of an incoming β-OMP substrate upon their release from the chaperone-bound state. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30018.map.gz emd_30018.map.gz | 92 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30018-v30.xml emd-30018-v30.xml emd-30018.xml emd-30018.xml | 16.8 KB 16.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30018.png emd_30018.png | 169.3 KB | ||
| Filedesc metadata |  emd-30018.cif.gz emd-30018.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30018 http://ftp.pdbj.org/pub/emdb/structures/EMD-30018 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30018 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30018 | HTTPS FTP |
-Related structure data
| Related structure data |  6lyuMC  6lyqC  6lyrC  6lysC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30018.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30018.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the BAM complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
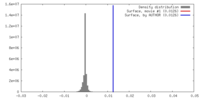
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : B-barrel assembly machinery (BAM) complex
| Entire | Name: B-barrel assembly machinery (BAM) complex |
|---|---|
| Components |
|
-Supramolecule #1: B-barrel assembly machinery (BAM) complex
| Supramolecule | Name: B-barrel assembly machinery (BAM) complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Outer membrane protein assembly factor BamA
| Macromolecule | Name: Outer membrane protein assembly factor BamA / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 90.643383 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAMKKLLIAS LLFSSATVYG AEGFVVKDIH FEGLQRVAVG AALLSMPVRT GDTVNDEDIS NTIRALFATG NFEDVRVLRD GDTLLVQVK ERPTIASITF SGNKSVKDDM LKQNLEASGV RVGESLDRTT IADIEKGLED FYYSVGKYSA SVKAVVTPLP R NRVDLKLV ...String: MAMKKLLIAS LLFSSATVYG AEGFVVKDIH FEGLQRVAVG AALLSMPVRT GDTVNDEDIS NTIRALFATG NFEDVRVLRD GDTLLVQVK ERPTIASITF SGNKSVKDDM LKQNLEASGV RVGESLDRTT IADIEKGLED FYYSVGKYSA SVKAVVTPLP R NRVDLKLV FQEGVSAEIQ QINIVGNHAF TTDELISHFQ LRDEVPWWNV VGDRKYQKQK LAGDLETLRS YYLDRGYARF NI DSTQVSL TPDKKGIYVT VNITEGDQYK LSGVEVSGNL AGHSAEIEQL TKIEPGELYN GTKVTKMEDD IKKLLGRYGY AYP RVQSMP EINDADKTVK LRVNVDAGNR FYVRKIRFEG NDTSKDAVLR REMRQMEGAW LGSDLVDQGK ERLNRLGFFE TVDT DTQRV PGSPDQVDVV YKVKERNTGS FNFGIGYGTE SGVSFQAGVQ QDNWLGTGYA VGINGTKNDY QTYAELSVTN PYFTV DGVS LGGRLFYNDF QADDADLSDY TNKSYGTDVT LGFPINEYNS LRAGLGYVHN SLSNMQPQVA MWRYLYSMGE HPSTSD QDN SFKTDDFTFN YGWTYNKLDR GYFPTDGSRV NLTGKVTIPG SDNEYYKVTL DTATYVPIDD DHKWVVLGRT RWGYGDG LG GKEMPFYENF YAGGSSTVRG FQSNTIGPKA VYFPHQASNY DPDYDYECAT QDGAKDLCKS DDAVGGNAMA VASLEFIT P TPFISDKYAN SVRTSFFWDM GTVWDTNWDS SQYSGYPDYS DPSNIRMSAG IALQWMSPLG PLVFSYAQPF KKYDGDKAE QFQFNIGKTW UniProtKB: Outer membrane protein assembly factor BamA |
-Macromolecule #2: Outer membrane protein assembly factor BamB
| Macromolecule | Name: Outer membrane protein assembly factor BamB / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.961086 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQLRKLLLPG LLSVTLLSGC SLFNSEEDVV KMSPLPTVEN QFTPTTAWST SVGSGIGNFY SNLHPALADN VVYAADRAGL VKALNADDG KEIWSVSLAE KDGWFSKEPA LLSGGVTVSG GHVYIGSEKA QVYALNTSDG TVAWQTKVAG EALSRPVVSD G LVLIHTSN ...String: MQLRKLLLPG LLSVTLLSGC SLFNSEEDVV KMSPLPTVEN QFTPTTAWST SVGSGIGNFY SNLHPALADN VVYAADRAGL VKALNADDG KEIWSVSLAE KDGWFSKEPA LLSGGVTVSG GHVYIGSEKA QVYALNTSDG TVAWQTKVAG EALSRPVVSD G LVLIHTSN GQLQALNEAD GAVKWTVNLD MPSLSLRGES APTTAFGAAV VGGDNGRVSA VLMEQGQMIW QQRISQATGS TE IDRLSDV DTTPVVVNGV VFALAYNGNL TALDLRSGQI MWKRELGSVN DFIVDGNRIY LVDQNDRVMA LTIDGGVTLW TQS DLLHRL LTSPVLYNGN LVVGDSEGYL HWINVEDGRF VAQQKVDSSG FQTEPVAADG KLLIQAKDGT VYSITRWSHP QFEK UniProtKB: Outer membrane protein assembly factor BamB |
-Macromolecule #3: Outer membrane protein assembly factor BamC
| Macromolecule | Name: Outer membrane protein assembly factor BamC / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.875277 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAYSVQKSRL AKVAGVSLVL LLAACSSDSR YKRQVSGDEA YLEAAPLAEL HAPAGMILPV TSGDYAIPVT NGSGAVGKAL DIRPPAQPL ALVSGARTQF TGDTASLLVE NGRGNTLWPQ VVSVLQAKNY TITQRDDAGQ TLTTDWVQWN RLDEDEQYRG R YQISVKPQ ...String: MAYSVQKSRL AKVAGVSLVL LLAACSSDSR YKRQVSGDEA YLEAAPLAEL HAPAGMILPV TSGDYAIPVT NGSGAVGKAL DIRPPAQPL ALVSGARTQF TGDTASLLVE NGRGNTLWPQ VVSVLQAKNY TITQRDDAGQ TLTTDWVQWN RLDEDEQYRG R YQISVKPQ GYQQAVTVKL LNLEQAGKPV ADAASMQRYS TEMMNVISAG LDKSATDAAN AAQNRASTTM DVQSAADDTG LP MLVVRGP FNVVWQRLPA ALEKVGMKVT DSTRSQGNMA VTYKPLSDSD WQELGASDPG LASGDYKLQV GDLDNRSSLQ FID PKGHTL TQSQNDALVA VFQAAFSK UniProtKB: Outer membrane protein assembly factor BamC |
-Macromolecule #4: Outer membrane protein assembly factor BamD
| Macromolecule | Name: Outer membrane protein assembly factor BamD / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.85835 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTRMKYLVAA ATLSLFLAGC SGSKEEVPDN PPNEIYATAQ QKLQDGNWRQ AITQLEALDN RYPFGPYSQQ VQLDLIYAYY KNADLPLAQ AAIDRFIRLN PTHPNIDYVM YMRGLTNMAL DDSALQGFFG VDRSDRDPQH ARAAFSDFSK LVRGYPNSQY T TDATKRLV ...String: MTRMKYLVAA ATLSLFLAGC SGSKEEVPDN PPNEIYATAQ QKLQDGNWRQ AITQLEALDN RYPFGPYSQQ VQLDLIYAYY KNADLPLAQ AAIDRFIRLN PTHPNIDYVM YMRGLTNMAL DDSALQGFFG VDRSDRDPQH ARAAFSDFSK LVRGYPNSQY T TDATKRLV FLKDRLAKYE YSVAEYYTER GAWVAVVNRV EGMLRDYPDT QATRDALPLM ENAYRQMQMN AQAEKVAKII AA NSSNT UniProtKB: Outer membrane protein assembly factor BamD |
-Macromolecule #5: Outer membrane protein assembly factor BamE
| Macromolecule | Name: Outer membrane protein assembly factor BamE / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.139857 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRCKTLTAAA AVLLMLTAGC STLERVVYRP DINQGNYLTA NDVSKIRVGM TQQQVAYALG TPLMSDPFGT NTWFYVFRQQ PGHEGVTQQ TLTLTFNSSG VLTNIDNKPA LSGNHHHHHH UniProtKB: Outer membrane protein assembly factor BamE |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)