+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2886 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-Molecular electron tomography of PfEMP1-IT4Var60) | |||||||||
 Map data Map data | Subtomogram average of 10 PfEMP1s | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Malaria / Rosetting / PfEMP1 | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 20.0 Å | |||||||||
 Authors Authors | Akhouri RR / Goel S / Furusho H / Skoglund U / Wahlgren M | |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2016 Journal: Cell Rep / Year: 2016Title: Architecture of Human IgM in Complex with P. falciparum Erythrocyte Membrane Protein 1. Authors: Reetesh Raj Akhouri / Suchi Goel / Hirotoshi Furusho / Ulf Skoglund / Mats Wahlgren /   Abstract: Plasmodium falciparum virulence is associated with sequestration of infected erythrocytes. Microvascular binding mediated by PfEMP1 in complex with non-immune immunoglobulin M (IgM) is common among ...Plasmodium falciparum virulence is associated with sequestration of infected erythrocytes. Microvascular binding mediated by PfEMP1 in complex with non-immune immunoglobulin M (IgM) is common among parasites that cause both severe childhood malaria and pregnancy-associated malaria. Here, we present cryo-molecular electron tomography structures of human IgM, PfEMP1 and their complex. Three-dimensional reconstructions of IgM reveal that it has a dome-like core, randomly oriented Fab2s units, and the overall shape of a turtle. PfEMP1 is a C- shaped molecule with a flexible N terminus followed by an arc-shaped backbone and a bulky C terminus that interacts with IgM. Our data demonstrate that the PfEMP1 binding pockets on IgM overlap with those of C1q, and the bulkiness of PfEMP1 limits the capacity of IgM to interact with PfEMP1. We suggest that P. falciparum exploits IgM to cluster PfEMP1 into an organized matrix to augment its affinity to host cell receptors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2886.map.gz emd_2886.map.gz | 403.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2886-v30.xml emd-2886-v30.xml emd-2886.xml emd-2886.xml | 9.3 KB 9.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2886.tif emd_2886.tif | 47.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2886 http://ftp.pdbj.org/pub/emdb/structures/EMD-2886 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2886 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2886 | HTTPS FTP |
-Related structure data
| Related structure data | 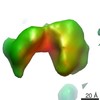 2882C  2883C  2884C  2885C  2887C  2888C  2889C  2890C  2891C  2892C  2893C  2894C  2895C  6554C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2886.map.gz / Format: CCP4 / Size: 450.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2886.map.gz / Format: CCP4 / Size: 450.2 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of 10 PfEMP1s | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.534 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ectodomain of PfEMP1-IT4Var60
| Entire | Name: ectodomain of PfEMP1-IT4Var60 |
|---|---|
| Components |
|
-Supramolecule #1000: ectodomain of PfEMP1-IT4Var60
| Supramolecule | Name: ectodomain of PfEMP1-IT4Var60 / type: sample / ID: 1000 / Details: the sample was monodisperse. / Oligomeric state: Monomer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 320 KDa / Theoretical: 280 KDa |
-Macromolecule #1: PfEMP1
| Macromolecule | Name: PfEMP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Strain: FCR3S1.2 / synonym: Malaria Parasite / Location in cell: Infected erythrocyte membrane |
| Molecular weight | Experimental: 320 KDa / Theoretical: 280 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 20mM Hepes, 200mM NaCl |
| Grid | Details: C-Flat (Copper grid with thin Carbon support) |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 78 K / Instrument: FEI VITROBOT MARK IV / Method: Blot for 4 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 78 K / Max: 100 K |
| Specialist optics | Energy filter - Name: FEI |
| Date | Oct 25, 2013 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Number real images: 281 / Average electron dose: 40 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 0.0015 µm / Nominal defocus min: 0.0007 µm / Nominal magnification: 37000 |
| Sample stage | Specimen holder: LN2 cooled / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Tilt series - Axis1 - Min angle: -70 ° / Tilt series - Axis1 - Max angle: 70 ° |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Software - Name: COMET / Number subtomograms used: 10 |
|---|---|
| CTF correction | Details: each frame |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















