+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Liver carboxylesterase 1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | carboxylesterase / human liver / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationcholesterol ester hydrolysis involved in cholesterol transport / methylumbelliferyl-acetate deacetylase / methylumbelliferyl-acetate deacetylase activity / regulation of bile acid secretion / sterol esterase / sterol ester esterase activity / medium-chain fatty acid metabolic process / carboxylesterase / Physiological factors / carboxylesterase activity ...cholesterol ester hydrolysis involved in cholesterol transport / methylumbelliferyl-acetate deacetylase / methylumbelliferyl-acetate deacetylase activity / regulation of bile acid secretion / sterol esterase / sterol ester esterase activity / medium-chain fatty acid metabolic process / carboxylesterase / Physiological factors / carboxylesterase activity / regulation of bile acid biosynthetic process / cellular response to cholesterol / reverse cholesterol transport / positive regulation of cholesterol metabolic process / Phase I - Functionalization of compounds / carboxylic ester hydrolase activity / cholesterol biosynthetic process / Aspirin ADME / negative regulation of cholesterol storage / positive regulation of cholesterol efflux / cellular response to low-density lipoprotein particle stimulus / Metabolism of Angiotensinogen to Angiotensins / cholesterol metabolic process / lipid catabolic process / epithelial cell differentiation / lipid droplet / cholesterol homeostasis / response to toxic substance / endoplasmic reticulum lumen / endoplasmic reticulum / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.67 Å | |||||||||
 Authors Authors | Zhang Z / Yu E | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2023 Journal: Cell Rep / Year: 2023Title: High-resolution structural-omics of human liver enzymes. Authors: Chih-Chia Su / Meinan Lyu / Zhemin Zhang / Masaru Miyagi / Wei Huang / Derek J Taylor / Edward W Yu /  Abstract: We applied raw human liver microsome lysate to a holey carbon grid and used cryo-electron microscopy (cryo-EM) to define its composition. From this sample we identified and simultaneously determined ...We applied raw human liver microsome lysate to a holey carbon grid and used cryo-electron microscopy (cryo-EM) to define its composition. From this sample we identified and simultaneously determined high-resolution structural information for ten unique human liver enzymes involved in diverse cellular processes. Notably, we determined the structure of the endoplasmic bifunctional protein H6PD, where the N- and C-terminal domains independently possess glucose-6-phosphate dehydrogenase and 6-phosphogluconolactonase enzymatic activity, respectively. We also obtained the structure of heterodimeric human GANAB, an ER glycoprotein quality-control machinery that contains a catalytic α subunit and a noncatalytic β subunit. In addition, we observed a decameric peroxidase, PRDX4, which directly contacts a disulfide isomerase-related protein, ERp46. Structural data suggest that several glycosylations, bound endogenous compounds, and ions associate with these human liver enzymes. These results highlight the importance of cryo-EM in facilitating the elucidation of human organ proteomics at the atomic level. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28465.map.gz emd_28465.map.gz | 97.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28465-v30.xml emd-28465-v30.xml emd-28465.xml emd-28465.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28465_fsc.xml emd_28465_fsc.xml | 9.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_28465.png emd_28465.png | 170.1 KB | ||
| Filedesc metadata |  emd-28465.cif.gz emd-28465.cif.gz | 5.4 KB | ||
| Others |  emd_28465_additional_1.map.gz emd_28465_additional_1.map.gz emd_28465_half_map_1.map.gz emd_28465_half_map_1.map.gz emd_28465_half_map_2.map.gz emd_28465_half_map_2.map.gz | 52 MB 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28465 http://ftp.pdbj.org/pub/emdb/structures/EMD-28465 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28465 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28465 | HTTPS FTP |
-Related structure data
| Related structure data |  8eorMC  7uzmC  8ekwC  8ekyC  8em2C  8emrC  8emsC  8emtC  8eneC  8eojC  23427 M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28465.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28465.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_28465_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_28465_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_28465_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Liver carboxylesterase 1
| Entire | Name: Liver carboxylesterase 1 |
|---|---|
| Components |
|
-Supramolecule #1: Liver carboxylesterase 1
| Supramolecule | Name: Liver carboxylesterase 1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Liver carboxylesterase 1
| Macromolecule | Name: Liver carboxylesterase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: carboxylesterase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 62.587773 KDa |
| Sequence | String: MWLRAFILAT LSASAAWGHP SSPPVVDTVH GKVLGKFVSL EGFAQPVAIF LGIPFAKPPL GPLRFTPPQP AEPWSFVKNA TSYPPMCTQ DPKAGQLLSE LFTNRKENIP LKLSEDCLYL NIYTPADLTK KNRLPVMVWI HGGGLMVGAA STYDGLALAA H ENVVVVTI ...String: MWLRAFILAT LSASAAWGHP SSPPVVDTVH GKVLGKFVSL EGFAQPVAIF LGIPFAKPPL GPLRFTPPQP AEPWSFVKNA TSYPPMCTQ DPKAGQLLSE LFTNRKENIP LKLSEDCLYL NIYTPADLTK KNRLPVMVWI HGGGLMVGAA STYDGLALAA H ENVVVVTI QYRLGIWGFF STGDEHSRGN WGHLDQVAAL RWVQDNIASF GGNPGSVTIF GESAGGESVS VLVLSPLAKN LF HRAISES GVALTSVLVK KGDVKPLAEQ IAITAGCKTT TSAVMVHCLR QKTEEELLET TLKMKFLSLD LQGDPRESQP LLG TVIDGM LLLKTPEELQ AERNFHTVPY MVGINKQEFG WLIPMQLMSY PLSEGQLDQK TAMSLLWKSY PLVCIAKELI PEAT EKYLG GTDDTVKKKD LFLDLIADVM FGVPSVIVAR NHRDAGAPTY MYEFQYRPSF SSDMKPKTVI GDHGDELFSV FGAPF LKEG ASEEEIRLSK MVMKFWANFA RNGNPNGEGL PHWPEYNQKE GYLQIGANTQ AAQKLKDKEV AFWTNLFAKK AVEKPP QTE HIEL UniProtKB: Liver carboxylesterase 1 |
-Macromolecule #3: ETHYL ACETATE
| Macromolecule | Name: ETHYL ACETATE / type: ligand / ID: 3 / Number of copies: 3 / Formula: EEE |
|---|---|
| Molecular weight | Theoretical: 88.105 Da |
| Chemical component information | 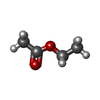 ChemComp-EEE: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 41.25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.291 µm / Nominal defocus min: 0.17 µm / Nominal magnification: 81000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)