+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24317 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of human ABCG1 E242Q complexed with ATP | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | sterol / lipids / ABC transporter / LIPID TRANSPORT | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationABC-type sterol transporter activity / cellular response to high density lipoprotein particle stimulus / intracellular cholesterol transport / glycoprotein transport / ABC transporters in lipid homeostasis / floppase activity / toxin transmembrane transporter activity / phospholipid efflux / phosphatidylcholine floppase activity / high-density lipoprotein particle remodeling ...ABC-type sterol transporter activity / cellular response to high density lipoprotein particle stimulus / intracellular cholesterol transport / glycoprotein transport / ABC transporters in lipid homeostasis / floppase activity / toxin transmembrane transporter activity / phospholipid efflux / phosphatidylcholine floppase activity / high-density lipoprotein particle remodeling / reverse cholesterol transport / phospholipid homeostasis / cholesterol transfer activity / low-density lipoprotein particle remodeling / xenobiotic detoxification by transmembrane export across the plasma membrane / HDL remodeling / cholesterol efflux / Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate / regulation of cholesterol metabolic process / positive regulation of amyloid-beta formation / cholesterol binding / amyloid precursor protein catabolic process / response to lipid / negative regulation of cholesterol storage / negative regulation of macrophage derived foam cell differentiation / positive regulation of cholesterol efflux / cholesterol metabolic process / NR1H3 & NR1H2 regulate gene expression linked to cholesterol transport and efflux / cholesterol homeostasis / positive regulation of protein secretion / ADP binding / positive regulation of cholesterol biosynthetic process / phospholipid binding / recycling endosome / transmembrane transport / endosome / protein heterodimerization activity / Golgi membrane / external side of plasma membrane / endoplasmic reticulum membrane / Golgi apparatus / protein homodimerization activity / ATP hydrolysis activity / mitochondrion / nucleoplasm / ATP binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
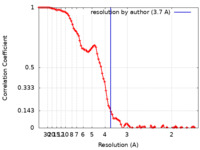
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | ||||||||||||
 Authors Authors | Sun Y / Li X | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2021 Journal: Proc Natl Acad Sci U S A / Year: 2021Title: Molecular basis of cholesterol efflux via ABCG subfamily transporters. Authors: Yingyuan Sun / Jin Wang / Tao Long / Xiaofeng Qi / Linda Donnelly / Nadia Elghobashi-Meinhardt / Leticia Esparza / Jonathan C Cohen / Xiao-Song Xie / Helen H Hobbs / Xiaochun Li /   Abstract: The ABCG1 homodimer (G1) and ABCG5-ABCG8 heterodimer (G5G8), two members of the adenosine triphosphate (ATP)-binding cassette (ABC) transporter G family, are required for maintenance of cellular ...The ABCG1 homodimer (G1) and ABCG5-ABCG8 heterodimer (G5G8), two members of the adenosine triphosphate (ATP)-binding cassette (ABC) transporter G family, are required for maintenance of cellular cholesterol levels. G5G8 mediates secretion of neutral sterols into bile and the gut lumen, whereas G1 transports cholesterol from macrophages to high-density lipoproteins (HDLs). The mechanisms used by G5G8 and G1 to recognize and export sterols remain unclear. Here, we report cryoelectron microscopy (cryo-EM) structures of human G5G8 in sterol-bound and human G1 in cholesterol- and ATP-bound states. Both transporters have a sterol-binding site that is accessible from the cytosolic leaflet. A second site is present midway through the transmembrane domains of G5G8. The Walker A motif of G8 adopts a unique conformation that accounts for the marked asymmetry in ATPase activities between the two nucleotide-binding sites of G5G8. These structures, along with functional validation studies, provide a mechanistic framework for understanding cholesterol efflux via ABC transporters. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24317.map.gz emd_24317.map.gz | 10.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24317-v30.xml emd-24317-v30.xml emd-24317.xml emd-24317.xml | 11.9 KB 11.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24317_fsc.xml emd_24317_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_24317.png emd_24317.png | 48.7 KB | ||
| Filedesc metadata |  emd-24317.cif.gz emd-24317.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24317 http://ftp.pdbj.org/pub/emdb/structures/EMD-24317 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24317 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24317 | HTTPS FTP |
-Related structure data
| Related structure data |  7r8eMC  7r87C  7r88C  7r89C  7r8aC  7r8bC  7r8cC  7r8dC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24317.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24317.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.833 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
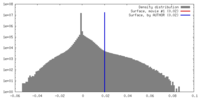
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ABCG1 transporter with cholesterol and ATP bound
| Entire | Name: ABCG1 transporter with cholesterol and ATP bound |
|---|---|
| Components |
|
-Supramolecule #1: ABCG1 transporter with cholesterol and ATP bound
| Supramolecule | Name: ABCG1 transporter with cholesterol and ATP bound / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: Isoform 4 of ATP-binding cassette sub-family G member 1
| Macromolecule | Name: Isoform 4 of ATP-binding cassette sub-family G member 1 type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO EC number: Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 74.228 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MACLMAAFSV GTAMNASSYS AEMTEPKSVC VSVDEVVSSN MEATETDLLN GHLKKVDNNL TEAQRFSSLP RRAAVNIEFR DLSYSVPEG PWWRKKGYKT LLKGISGKFN SGELVAIMGP SGAGKSTLMN ILAGYRETGM KGAVLINGLP RDLRCFRKVS C YIMQDDML ...String: MACLMAAFSV GTAMNASSYS AEMTEPKSVC VSVDEVVSSN MEATETDLLN GHLKKVDNNL TEAQRFSSLP RRAAVNIEFR DLSYSVPEG PWWRKKGYKT LLKGISGKFN SGELVAIMGP SGAGKSTLMN ILAGYRETGM KGAVLINGLP RDLRCFRKVS C YIMQDDML LPHLTVQEAM MVSAHLKLQE KDEGRREMVK EILTALGLLS CANTRTGSLS GGQRKRLAIA LELVNNPPVM FF DQPTSGL DSASCFQVVS LMKGLAQGGR SIICTIHQPS AKLFELFDQL YVLSQGQCVY RGKVCNLVPY LRDLGLNCPT YHN PADFVM EVASGEYGDQ NSRLVRAVRE GMCDSDHKRD LGGDAEVNPF LWHRPSEEDS SSMEGCHSFS ASCLTQFCIL FKRT FLSIM RDSVLTHLRI TSHIGIGLLI GLLYLGIGNE AKKVLSNSGF LFFSMLFLMF AALMPTVLTF PLEMGVFLRE HLNYW YSLK AYYLAKTMAD VPFQIMFPVA YCSIVYWMTS QPSDAVRFVL FAALGTMTSL VAQSLGLLIG AASTSLQVAT FVGPVT AIP VLLFSGFFVS FDTIPTYLQW MSYISYVRYG FEGVILSIYG LDREDLHCDI DETCHFQKSE AILRELDVEN AKLYLDF IV LGIFFISLRL IAYFVLRYKI RAER UniProtKB: ATP-binding cassette sub-family G member 1 |
-Macromolecule #2: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 2 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #4: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 4 / Number of copies: 2 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
| Details | Added 8mM ATP, Magnesium and 0.2mg/ml cholesterol in ethanol |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 4900 / Average exposure time: 1.8 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)