+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24311 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of human ABCG5-I529W/ABCG8-WT | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | sterol / lipids / ABC transporter / LIPID TRANSPORT / LIPID TRANSPORT-IMMUNE SYSTEM complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of intestinal phytosterol absorption / negative regulation of intestinal cholesterol absorption / Defective ABCG8 causes GBD4 and sitosterolemia / Defective ABCG5 causes sitosterolemia / sterol transport / intestinal cholesterol absorption / ABC transporters in lipid homeostasis / phospholipid transport / cholesterol transfer activity / cholesterol efflux ...negative regulation of intestinal phytosterol absorption / negative regulation of intestinal cholesterol absorption / Defective ABCG8 causes GBD4 and sitosterolemia / Defective ABCG5 causes sitosterolemia / sterol transport / intestinal cholesterol absorption / ABC transporters in lipid homeostasis / phospholipid transport / cholesterol transfer activity / cholesterol efflux / Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate / triglyceride homeostasis / response to muscle activity / response to ionizing radiation / ATPase-coupled transmembrane transporter activity / ABC-type transporter activity / NR1H3 & NR1H2 regulate gene expression linked to cholesterol transport and efflux / response to nutrient / ATP-binding cassette (ABC) transporter complex / cholesterol homeostasis / transmembrane transport / signaling receptor complex / apical plasma membrane / response to xenobiotic stimulus / protein heterodimerization activity / ATP hydrolysis activity / ATP binding / metal ion binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
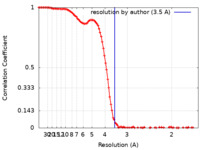
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||
 Authors Authors | Sun Y / Li X | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2021 Journal: Proc Natl Acad Sci U S A / Year: 2021Title: Molecular basis of cholesterol efflux via ABCG subfamily transporters. Authors: Yingyuan Sun / Jin Wang / Tao Long / Xiaofeng Qi / Linda Donnelly / Nadia Elghobashi-Meinhardt / Leticia Esparza / Jonathan C Cohen / Xiao-Song Xie / Helen H Hobbs / Xiaochun Li /   Abstract: The ABCG1 homodimer (G1) and ABCG5-ABCG8 heterodimer (G5G8), two members of the adenosine triphosphate (ATP)-binding cassette (ABC) transporter G family, are required for maintenance of cellular ...The ABCG1 homodimer (G1) and ABCG5-ABCG8 heterodimer (G5G8), two members of the adenosine triphosphate (ATP)-binding cassette (ABC) transporter G family, are required for maintenance of cellular cholesterol levels. G5G8 mediates secretion of neutral sterols into bile and the gut lumen, whereas G1 transports cholesterol from macrophages to high-density lipoproteins (HDLs). The mechanisms used by G5G8 and G1 to recognize and export sterols remain unclear. Here, we report cryoelectron microscopy (cryo-EM) structures of human G5G8 in sterol-bound and human G1 in cholesterol- and ATP-bound states. Both transporters have a sterol-binding site that is accessible from the cytosolic leaflet. A second site is present midway through the transmembrane domains of G5G8. The Walker A motif of G8 adopts a unique conformation that accounts for the marked asymmetry in ATPase activities between the two nucleotide-binding sites of G5G8. These structures, along with functional validation studies, provide a mechanistic framework for understanding cholesterol efflux via ABC transporters. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24311.map.gz emd_24311.map.gz | 11.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24311-v30.xml emd-24311-v30.xml emd-24311.xml emd-24311.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24311_fsc.xml emd_24311_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_24311.png emd_24311.png | 129.2 KB | ||
| Filedesc metadata |  emd-24311.cif.gz emd-24311.cif.gz | 7.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24311 http://ftp.pdbj.org/pub/emdb/structures/EMD-24311 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24311 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24311 | HTTPS FTP |
-Related structure data
| Related structure data |  7r88MC  7r87C  7r89C  7r8aC  7r8bC  7r8cC  7r8dC  7r8eC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24311.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24311.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.833 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
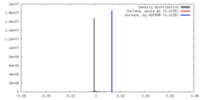
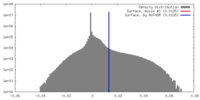
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ABCG5-I529W/ABCG8 transporter with Fab 2C7 bound
| Entire | Name: ABCG5-I529W/ABCG8 transporter with Fab 2C7 bound |
|---|---|
| Components |
|
-Supramolecule #1: ABCG5-I529W/ABCG8 transporter with Fab 2C7 bound
| Supramolecule | Name: ABCG5-I529W/ABCG8 transporter with Fab 2C7 bound / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 150 KDa |
-Supramolecule #2: ABCG5-I529W/ABCG8 transporter
| Supramolecule | Name: ABCG5-I529W/ABCG8 transporter / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Fab 2C7
| Supramolecule | Name: Fab 2C7 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: ATP-binding cassette sub-family G member 5
| Macromolecule | Name: ATP-binding cassette sub-family G member 5 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 74.510438 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGDLSSLTPG GSMGLQVNRG SQSSLEGAPA TAPEPHSLGI LHASYSVSHR VRPWWDITSC RQQWTRQILK DVSLYVESGQ IMCILGSSG SGKTTLLDAM SGRLGRAGTF LGEVYVNGRA LRREQFQDCF SYVLQSDTLL SSLTVRETLH YTALLAIRRG N PGSFQKKV ...String: MGDLSSLTPG GSMGLQVNRG SQSSLEGAPA TAPEPHSLGI LHASYSVSHR VRPWWDITSC RQQWTRQILK DVSLYVESGQ IMCILGSSG SGKTTLLDAM SGRLGRAGTF LGEVYVNGRA LRREQFQDCF SYVLQSDTLL SSLTVRETLH YTALLAIRRG N PGSFQKKV EAVMAELSLS HVADRLIGNY SLGGISTGER RRVSIAAQLL QDPKVMLFDE PTTGLDCMTA NQIVVLLVEL AR RNRIVVL TIHQPRSELF QLFDKIAILS FGELIFCGTP AEMLDFFNDC GYPCPEHSNP FDFYMDLTSV DTQSKEREIE TSK RVQMIE SAYKKSAICH KTLKNIERMK HLKTLPMVPF KTKDSPGVFS KLGVLLRRVT RNLVRNKLAV ITRLLQNLIM GLFL LFFVL RVRSNVLKGA IQDRVGLLYQ FVGATPYTGM LNAVNLFPVL RAVSDQESQD GLYQKWQMML AYALHVLPFS VVATM IFSS VCYWTLGLHP EVARFGYFSA ALLAPHLIGE FLTLVLLGIV QNPNWVNSVV ALLSIAGVLV GSGFLRNIQE MPIPFK IIS YFTFQKYCSE ILVVNEFYGL NFTCGSSNVS VTTNPMCAFT QGIQFIEKTC PGATSRFTMN FLILYSFIPA LVILGIV VF KIRDHLISRG SHHHHHHGHH HHHH UniProtKB: ATP-binding cassette sub-family G member 5 |
-Macromolecule #2: ATP-binding cassette sub-family G member 8
| Macromolecule | Name: ATP-binding cassette sub-family G member 8 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO EC number: Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 80.365828 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAGKAAEERG LPKGATPQDT SGLQDRLFSS ESDNSLYFTY SGQPNTLEVR DLNYQVDLAS QVPWFEQLAQ FKMPWTSPSC QNSCELGIQ NLSFKVRSGQ MLAIIGSSGC GRASLLDVIT GRGHGGKIKS GQIWINGQPS SPQLVRKCVA HVRQHNQLLP N LTVRETLA ...String: MAGKAAEERG LPKGATPQDT SGLQDRLFSS ESDNSLYFTY SGQPNTLEVR DLNYQVDLAS QVPWFEQLAQ FKMPWTSPSC QNSCELGIQ NLSFKVRSGQ MLAIIGSSGC GRASLLDVIT GRGHGGKIKS GQIWINGQPS SPQLVRKCVA HVRQHNQLLP N LTVRETLA FIAQMRLPRT FSQAQRDKRV EDVIAELRLR QCADTRVGNM YVRGLSGGER RRVSIGVQLL WNPGILILDE PT SGLDSFT AHNLVKTLSR LAKGNRLVLI SLHQPRSDIF RLFDLVLLMT SGTPIYLGAA QHMVQYFTAI GYPCPRYSNP ADF YVDLTS IDRRSREQEL ATREKAQSLA ALFLEKVRDL DDFLWKAETK DLDEDTCVES SVTPLDTNCL PSPTKMPGAV QQFT TLIRR QISNDFRDLP TLLIHGAEAC LMSMTIGFLY FGHGSIQLSF MDTAALLFMI GALIPFNVIL DVISKCYSER AMLYY ELED GLYTTGPYFF AKILGELPEH CAYIIIYGMP TYWLANLRPG LQPFLLHFLL VWLVVFCCRI MALAAAALLP TFHMAS FFS NALYNSFYLA GGFMINLSSL WTVPAWISKV SFLRWCFEGL MKIQFSRRTY KMPLGNLTIA VSGDKILSVM ELDSYPL YA IYLIVIGLSG GFMVLYYVSL RFIKQKPSQD WASNSLEVLF QGPNVDSKRR WKKNFIAVSA ANRFKKISSS GAL UniProtKB: ATP-binding cassette sub-family G member 8 |
-Macromolecule #3: 2C7 Fab heavy chain
| Macromolecule | Name: 2C7 Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.37967 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGWSCIILFL VATATGVHSE VKLVESGGGL VQPGGSLRLS CATSGFTFSE FFMEWVRQPP GKRLEWVAVS RNEANDYTTD YSASVKGRF IVSRDTSQNI LYLQMNALRA EDTAIYYCAR DAWMGFDYWG QGTTVTVSSA STKGPSVFPL APSSKSTSGG T AALGCLVK ...String: MGWSCIILFL VATATGVHSE VKLVESGGGL VQPGGSLRLS CATSGFTFSE FFMEWVRQPP GKRLEWVAVS RNEANDYTTD YSASVKGRF IVSRDTSQNI LYLQMNALRA EDTAIYYCAR DAWMGFDYWG QGTTVTVSSA STKGPSVFPL APSSKSTSGG T AALGCLVK DYFPEPVTVS WNSGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YICNVNHKPS NTKVDKRVEP KS CDKTH |
-Macromolecule #4: 2C7 Fab light chain
| Macromolecule | Name: 2C7 Fab light chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.605615 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGWSCIILFL VATARTGVHS DIQMTQSPSS LSASLGERVS LTCRASQEIS GYLSWLQQKP DGTIQRLIYA AFSLDSGVPK RFSGSRSGS DYSLTISSLE SEDLAHYYCL QYASYPCTFG GGTKLEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF Y PREAKVQW ...String: MGWSCIILFL VATARTGVHS DIQMTQSPSS LSASLGERVS LTCRASQEIS GYLSWLQQKP DGTIQRLIYA AFSLDSGVPK RFSGSRSGS DYSLTISSLE SEDLAHYYCL QYASYPCTFG GGTKLEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF Y PREAKVQW KVDNALQSGN SQESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTH QGLSSPVTKS FNRGEC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 4900 / Average exposure time: 1.8 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)