[English] 日本語
 Yorodumi
Yorodumi- EMDB-22852: Ebola virus GP (mucin deleted, Makona strain) bound to antibody F... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22852 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ebola virus GP (mucin deleted, Makona strain) bound to antibody Fab EBOV-237 and EBOV-515 | |||||||||
 Map data Map data | Ebola virus GP (mucin deleted, Makona strain) bound to antibody Fab EBOV-237 and EBOV-515 | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationmembrane => GO:0016020 / clathrin-dependent endocytosis of virus by host cell / symbiont-mediated-mediated suppression of host tetherin activity / entry receptor-mediated virion attachment to host cell / symbiont-mediated suppression of host innate immune response / fusion of virus membrane with host endosome membrane / viral envelope / lipid binding / host cell plasma membrane / virion membrane ...membrane => GO:0016020 / clathrin-dependent endocytosis of virus by host cell / symbiont-mediated-mediated suppression of host tetherin activity / entry receptor-mediated virion attachment to host cell / symbiont-mediated suppression of host innate immune response / fusion of virus membrane with host endosome membrane / viral envelope / lipid binding / host cell plasma membrane / virion membrane / extracellular region / membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
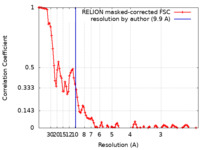
| Method | single particle reconstruction / cryo EM / Resolution: 9.9 Å | |||||||||
 Authors Authors | Murin CD / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2021 Journal: Cell Rep / Year: 2021Title: Convergence of a common solution for broad ebolavirus neutralization by glycan cap-directed human antibodies. Authors: Charles D Murin / Pavlo Gilchuk / Philipp A Ilinykh / Kai Huang / Natalia Kuzmina / Xiaoli Shen / Jessica F Bruhn / Aubrey L Bryan / Edgar Davidson / Benjamin J Doranz / Lauren E Williamson ...Authors: Charles D Murin / Pavlo Gilchuk / Philipp A Ilinykh / Kai Huang / Natalia Kuzmina / Xiaoli Shen / Jessica F Bruhn / Aubrey L Bryan / Edgar Davidson / Benjamin J Doranz / Lauren E Williamson / Jeffrey Copps / Tanwee Alkutkar / Andrew I Flyak / Alexander Bukreyev / James E Crowe / Andrew B Ward /  Abstract: Antibodies that target the glycan cap epitope on the ebolavirus glycoprotein (GP) are common in the adaptive response of survivors. A subset is known to be broadly neutralizing, but the details of ...Antibodies that target the glycan cap epitope on the ebolavirus glycoprotein (GP) are common in the adaptive response of survivors. A subset is known to be broadly neutralizing, but the details of their epitopes and basis for neutralization are not well understood. Here, we present cryoelectron microscopy (cryo-EM) structures of diverse glycan cap antibodies that variably synergize with GP base-binding antibodies. These structures describe a conserved site of vulnerability that anchors the mucin-like domains (MLDs) to the glycan cap, which we call the MLD anchor and cradle. Antibodies that bind to the MLD cradle share common features, including use of IGHV1-69 and IGHJ6 germline genes, which exploit hydrophobic residues and form β-hairpin structures to mimic the MLD anchor, disrupt MLD attachment, destabilize GP quaternary structure, and block cleavage events required for receptor binding. Our results provide a molecular basis for ebolavirus neutralization by broadly reactive glycan cap antibodies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22852.map.gz emd_22852.map.gz | 5.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22852-v30.xml emd-22852-v30.xml emd-22852.xml emd-22852.xml | 14.2 KB 14.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22852_fsc.xml emd_22852_fsc.xml | 10.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_22852.png emd_22852.png | 46.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22852 http://ftp.pdbj.org/pub/emdb/structures/EMD-22852 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22852 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22852 | HTTPS FTP |
-Validation report
| Summary document |  emd_22852_validation.pdf.gz emd_22852_validation.pdf.gz | 326.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22852_full_validation.pdf.gz emd_22852_full_validation.pdf.gz | 325.7 KB | Display | |
| Data in XML |  emd_22852_validation.xml.gz emd_22852_validation.xml.gz | 11.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22852 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22852 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22852 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22852 | HTTPS FTP |
-Related structure data
| Related structure data |  7kejC  7kewC  7kexC  7kf9C  7kfbC  7kfeC  7kfgC  7kfhC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22852.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22852.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ebola virus GP (mucin deleted, Makona strain) bound to antibody Fab EBOV-237 and EBOV-515 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ebola virus GP (mucin deleted, Makona strain) bound to antibody F...
| Entire | Name: Ebola virus GP (mucin deleted, Makona strain) bound to antibody Fab EBOV-237 and EBOV-515 |
|---|---|
| Components |
|
-Supramolecule #1: Ebola virus GP (mucin deleted, Makona strain) bound to antibody F...
| Supramolecule | Name: Ebola virus GP (mucin deleted, Makona strain) bound to antibody Fab EBOV-237 and EBOV-515 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Parts of complex expressed and purified recombinantly, complexed and purified by gel filtration |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: Antibody Fab EBOV-237
| Supramolecule | Name: Antibody Fab EBOV-237 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Ebola virus GP (mucin deleted, Makona strain)
| Supramolecule | Name: Ebola virus GP (mucin deleted, Makona strain) / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ebola virus GP1 (mucin deleted, Makona strain)
| Macromolecule | Name: Ebola virus GP1 (mucin deleted, Makona strain) / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDAMKRGLCC VLLLCGAVFV SPSQEIHARF RRGARSIPLG VIHNSTLQVS DVDKLVCRDK LSSTNQLRSV GLNLEGNGVA TDVPSVTKRW GFRSGVPPKV VNYEAGEWAE NCYNLEIKKP DGSECLPAAP DGIRGFPRCR YVHKVSGTGP CAGDFAFHKE GAFFLYDRLA ...String: MDAMKRGLCC VLLLCGAVFV SPSQEIHARF RRGARSIPLG VIHNSTLQVS DVDKLVCRDK LSSTNQLRSV GLNLEGNGVA TDVPSVTKRW GFRSGVPPKV VNYEAGEWAE NCYNLEIKKP DGSECLPAAP DGIRGFPRCR YVHKVSGTGP CAGDFAFHKE GAFFLYDRLA STVIYRGTTF AEGVVAFLIL PQAKKDFFSS HPLREPVNAT EDPSSGYYST TIRYQATGFG TNETEYLFEV DNLTYVQLES RFTPQFLLQL NETIYASGKR SNTTGKLIWK VNPEIDTTIG EWAFWETKKN LTRKIRSEEL SFT |
-Macromolecule #2: Ebola virus GP2 (Makona strain)
| Macromolecule | Name: Ebola virus GP2 (Makona strain) / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NNNTHHQDTG EESASSGKLG LITNTIAGVA GLITGGRRTR REVIVNAQPK CNPNLHYWTT QDEGAAIGLA WIPYFGPAAE GIYTEGLMHN QDGLICGLRQ LANETTQALQ LFLRATTELR TFSILNRKAI DFLLQRWGGT CHILGPDCCI EPHDWTKNIT DKIDQIIHDD ...String: NNNTHHQDTG EESASSGKLG LITNTIAGVA GLITGGRRTR REVIVNAQPK CNPNLHYWTT QDEGAAIGLA WIPYFGPAAE GIYTEGLMHN QDGLICGLRQ LANETTQALQ LFLRATTELR TFSILNRKAI DFLLQRWGGT CHILGPDCCI EPHDWTKNIT DKIDQIIHDD DDKAGWSHPQ FEKGGGSGGG SGGGSWSHPQ FEK |
-Macromolecule #3: Antibody Fab EBOV-237 heavy chain
| Macromolecule | Name: Antibody Fab EBOV-237 heavy chain / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPG LVKPSGTLSL TCAVSGGSIS NGIWWSWVRQ PPGKGLEWIG EIYHSGSTNY SPSLKSRVTI SLDKSKNQFS LKLNSVTAAD TAVYYCARAP SQFDFSDWLV PQGAMFRGHY IDYWGQGTLV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP ...String: QVQLQESGPG LVKPSGTLSL TCAVSGGSIS NGIWWSWVRQ PPGKGLEWIG EIYHSGSTNY SPSLKSRVTI SLDKSKNQFS LKLNSVTAAD TAVYYCARAP SQFDFSDWLV PQGAMFRGHY IDYWGQGTLV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP VTVSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HKPSNTKVDK RVEPKSCD |
-Macromolecule #4: Antibody Fab EBOV-237 light chain
| Macromolecule | Name: Antibody Fab EBOV-237 light chain / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIVMTQSPDS LAVSLGERAT INCKSSQSVL YNSNNKNYLA WYQQKPGQPP KLLIYWASTR ESGVPDRFSG SGSGTDFTLA ISSLRAEDVA VYYCQQYYSA PYTFGQGTKM EIKRTVAAPS VFIFPPSDEQ LKSGTASVVC LLNNFYPREA KVQWKVDNAL QSGNSQESVT ...String: DIVMTQSPDS LAVSLGERAT INCKSSQSVL YNSNNKNYLA WYQQKPGQPP KLLIYWASTR ESGVPDRFSG SGSGTDFTLA ISSLRAEDVA VYYCQQYYSA PYTFGQGTKM EIKRTVAAPS VFIFPPSDEQ LKSGTASVVC LLNNFYPREA KVQWKVDNAL QSGNSQESVT EQDSKDSTYS LSSTLTLSKA DYEKHKVYAC EVTHQGLSSP VTKSFNRGEC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)