[English] 日本語
 Yorodumi
Yorodumi- EMDB-21580: Single-Particle Cryo-EM Structure of Arabinofuranosyltransferase ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21580 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Single-Particle Cryo-EM Structure of Arabinofuranosyltransferase AftD from Mycobacteria | |||||||||||||||||||||
 Map data Map data | Sharpened map | |||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | Glycosyltransferase / lipomannan / lipoarabinomannan / arabinofuranose / membrane protein / nanodisc / single-particle cryo-electron microscopy / acyl carrier protein | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationlipid A biosynthetic process / lipid biosynthetic process / phosphopantetheine binding / acyl binding / membrane => GO:0016020 / acyl carrier activity / fatty acid biosynthetic process / transferase activity / response to xenobiotic stimulus / lipid binding ...lipid A biosynthetic process / lipid biosynthetic process / phosphopantetheine binding / acyl binding / membrane => GO:0016020 / acyl carrier activity / fatty acid biosynthetic process / transferase activity / response to xenobiotic stimulus / lipid binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||||||||||||||
| Biological species |   Mycobacteroides abscessus subsp. abscessus (bacteria) Mycobacteroides abscessus subsp. abscessus (bacteria) | |||||||||||||||||||||
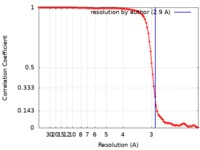
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||||||||||||||
 Authors Authors | Tan YZ / Zhang L | |||||||||||||||||||||
| Funding support |  United States, 6 items United States, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2020 Journal: Mol Cell / Year: 2020Title: Cryo-EM Structures and Regulation of Arabinofuranosyltransferase AftD from Mycobacteria. Authors: Yong Zi Tan / Lei Zhang / José Rodrigues / Ruixiang Blake Zheng / Sabrina I Giacometti / Ana L Rosário / Brian Kloss / Venkata P Dandey / Hui Wei / Richard Brunton / Ashleigh M Raczkowski ...Authors: Yong Zi Tan / Lei Zhang / José Rodrigues / Ruixiang Blake Zheng / Sabrina I Giacometti / Ana L Rosário / Brian Kloss / Venkata P Dandey / Hui Wei / Richard Brunton / Ashleigh M Raczkowski / Diogo Athayde / Maria João Catalão / Madalena Pimentel / Oliver B Clarke / Todd L Lowary / Margarida Archer / Michael Niederweis / Clinton S Potter / Bridget Carragher / Filippo Mancia /     Abstract: Mycobacterium tuberculosis causes tuberculosis, a disease that kills over 1 million people each year. Its cell envelope is a common antibiotic target and has a unique structure due, in part, to two ...Mycobacterium tuberculosis causes tuberculosis, a disease that kills over 1 million people each year. Its cell envelope is a common antibiotic target and has a unique structure due, in part, to two lipidated polysaccharides-arabinogalactan and lipoarabinomannan. Arabinofuranosyltransferase D (AftD) is an essential enzyme involved in assembling these glycolipids. We present the 2.9-Å resolution structure of M. abscessus AftD, determined by single-particle cryo-electron microscopy. AftD has a conserved GT-C glycosyltransferase fold and three carbohydrate-binding modules. Glycan array analysis shows that AftD binds complex arabinose glycans. Additionally, AftD is non-covalently complexed with an acyl carrier protein (ACP). 3.4- and 3.5-Å structures of a mutant with impaired ACP binding reveal a conformational change, suggesting that ACP may regulate AftD function. Mutagenesis experiments using a conditional knockout constructed in M. smegmatis confirm the essentiality of the putative active site and the ACP binding for AftD function. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21580.map.gz emd_21580.map.gz | 84.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21580-v30.xml emd-21580-v30.xml emd-21580.xml emd-21580.xml | 54.8 KB 54.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21580_fsc.xml emd_21580_fsc.xml | 10.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_21580.png emd_21580.png | 131.2 KB | ||
| Masks |  emd_21580_msk_1.map emd_21580_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-21580.cif.gz emd-21580.cif.gz | 9.7 KB | ||
| Others |  emd_21580_additional_1.map.gz emd_21580_additional_1.map.gz emd_21580_additional_10.map.gz emd_21580_additional_10.map.gz emd_21580_additional_11.map.gz emd_21580_additional_11.map.gz emd_21580_additional_2.map.gz emd_21580_additional_2.map.gz emd_21580_additional_3.map.gz emd_21580_additional_3.map.gz emd_21580_additional_4.map.gz emd_21580_additional_4.map.gz emd_21580_additional_5.map.gz emd_21580_additional_5.map.gz emd_21580_additional_6.map.gz emd_21580_additional_6.map.gz emd_21580_additional_7.map.gz emd_21580_additional_7.map.gz emd_21580_additional_8.map.gz emd_21580_additional_8.map.gz emd_21580_additional_9.map.gz emd_21580_additional_9.map.gz emd_21580_half_map_1.map.gz emd_21580_half_map_1.map.gz emd_21580_half_map_2.map.gz emd_21580_half_map_2.map.gz | 84.6 MB 14.8 MB 16.5 MB 84.6 MB 18.7 MB 16.2 MB 182.4 KB 84.6 MB 14.7 MB 172.7 KB 14.8 MB 14.7 MB 14.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21580 http://ftp.pdbj.org/pub/emdb/structures/EMD-21580 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21580 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21580 | HTTPS FTP |
-Related structure data
| Related structure data |  6w98MC  6wbxC  6wbyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10399 (Title: Single-Particle Cryo-EM of Arabinofuranosyltransferase AftD from Mycobacteria, Wild-type EMPIAR-10399 (Title: Single-Particle Cryo-EM of Arabinofuranosyltransferase AftD from Mycobacteria, Wild-typeData size: 2.7 TB Data #1: Unaligned and compressed multi-frame movies [micrographs - multiframe] Data #2: Aligned and dose-weighted micrographs [micrographs - single frame] Data #3: Final Particle Stacks with Refined Euler Angles and Shifts (Overall and Focused Refined on CBM3) [picked particles - single frame - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21580.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21580.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
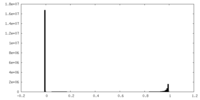
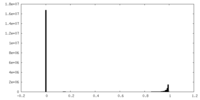
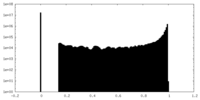
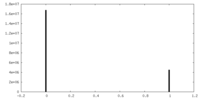
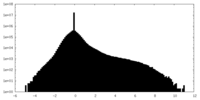
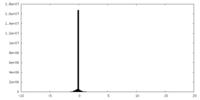
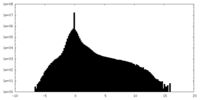
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0605 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
+Mask #1
+Additional map: Raw, unsharpened map
+Additional map: CBM3 focus classification - Half map 2
+Additional map: CBM3 focus classification - 3DFSC
+Additional map: Sharpened composite map of AftD WT with CBM3 focus classification
+Additional map: 3DFSC
+Additional map: 3DFSC Thresholded
+Additional map: 3DFSC Thresholded Binarized
+Additional map: CBM3 focus classification - Sharpened map
+Additional map: CBM3 focus classification - Raw, unsharpened map
+Additional map: CBM3 focus classification - Mask
+Additional map: CBM3 focus classification - Half map 1
+Half map: Half map 1
+Half map: Half map 2
- Sample components
Sample components
-Entire : Wild-type Mycobacterial Arabinofuranosyltransferase AftD Complexe...
| Entire | Name: Wild-type Mycobacterial Arabinofuranosyltransferase AftD Complexed with Acyl Carrier Protein |
|---|---|
| Components |
|
-Supramolecule #1: Wild-type Mycobacterial Arabinofuranosyltransferase AftD Complexe...
| Supramolecule | Name: Wild-type Mycobacterial Arabinofuranosyltransferase AftD Complexed with Acyl Carrier Protein type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Molecular weight | Theoretical: 50 KDa |
-Supramolecule #3: Acyl carrier protein
| Supramolecule | Name: Acyl carrier protein / type: organelle_or_cellular_component / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: F5/8 type C domain-containing protein
| Supramolecule | Name: F5/8 type C domain-containing protein / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Mycobacteroides abscessus subsp. abscessus (bacteria) Mycobacteroides abscessus subsp. abscessus (bacteria) |
-Macromolecule #1: F5/8 type C domain-containing protein
| Macromolecule | Name: F5/8 type C domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycobacteroides abscessus subsp. abscessus (bacteria) Mycobacteroides abscessus subsp. abscessus (bacteria) |
| Molecular weight | Theoretical: 149.584281 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SYVMTYRLDS SALSRRWLAV AAAVSLLLTF SQSPGQISPD TKLDLAINPL RFAARALNLW SSDLPFGQAQ NQAYGYLFPH GAFFSLGHL LGVPAWVTQR LWWALLIVAG FWGLIRVAEA LGIGTRGSRI IAAVAFALSP RVLTTLGAIS SETLPMMLAP W VLLPLILT ...String: SYVMTYRLDS SALSRRWLAV AAAVSLLLTF SQSPGQISPD TKLDLAINPL RFAARALNLW SSDLPFGQAQ NQAYGYLFPH GAFFSLGHL LGVPAWVTQR LWWALLIVAG FWGLIRVAEA LGIGTRGSRI IAAVAFALSP RVLTTLGAIS SETLPMMLAP W VLLPLILT FQGRMSPRRA AALSAVAVAL MGAVNAVATA LACGVAVIWW LAHRPNRTWW RFTAWWIPCL ALASTWWIVA LL IFGKISP KFLDFIESSG VTTQWTSLTE VLRGTDSWTP FVAPTATAGS SLVTQSAMVI ATTMLAAAGM AGLAMRGMPA RGR LVAVLL IGLVLLTAGY TGALGSPIAQ QIQFFLDDGG TPLRNVHKLE PLIRLPLILG LAHALSRIPL PASVPVRQWL SALA RPERN RAVAFAIVLL VALAASTSLA WTGRLVPRGG FDAIPGYWND TAHWLADHDT GGRALVVPGA PFAIQTWGLT RDEPL QALG QTPWGVRDSI PLTPPETIRA IDSVQQLFAA GRPSDGLADT LREQGISYLV VRNDLDPDTS RSARPILVHH TIEGSP GLT KVAQFGDPVG AGAVEGFVAD SDLRPQYPAV EIYAVGANDH DGEPYFTDID TMPRVAGGPE ALLRLNERRR QLNEPPL GP SLLATDAAQA GLRPGPAVVT DTPLARETDY GRVDDHSSAI RAPGDKRRTF NRVPDYPATG VPLVNGSWTG GTITASSS A SDSTALPNVA PGTSTAAAID RDNATSWVSS SLEAALGQWI RIDLDRPITN AILTVTPSAT ALGAQVRRLE VETDNGTTS VRFDEPGQPL NIALRPGETT WVKVTATGTD DGTSGVQFGV TELSLTQYDA AGFAHTVDLR HSATVPPPPA GDNPLGWDLG SPLQGRSGC APSPQRLRCA ATLSLAPEEP GTFIRTLTVP QPVSLTPRLW VRARPGPQLR DLIQQPGTTV ATGDSDVIDP Q GSSYAATD GDPGTVWTAP QDSVQRLHLP SLVIKLPKPT AIGAIRLRPS RTEVPAHPKQ VAINLGDGPQ LRSIDPKADV TE LALHPSI TDTITVTVTD WTDIIDRTAL GFDQLKPPGI AEVIALDADH RPIAPADNAA NSKRKITIGC NRGPILALAG RFV PMSITA TVRELLDGTV IQATPCDTSP IATGAGIQDV TVNPSQQFIV DGVQLTAAAT EPASATMTVA PKGAWGPDRR EVTA EPSAH ERVLAVPESI NPGWAARDAQ GHLLTPVRVN GWQQGWVLPA GDGGKITLTF GLNTWYRAGL FGGLALLPIL ACLAL LPAR GRTTLPPVAP WCAGPAAGVA VLAALTAISG ISGMAVGLAA LAFKVWTRWP LRAVTAAGVY LAGGSLLLAG AALSRH PWR SVGGYTGHSW WIQLLALISV ASVALAAVRL PSRRCWKRRS ASREGDSTSA UniProtKB: F5/8 type C domain-containing protein |
-Macromolecule #2: Acyl carrier protein
| Macromolecule | Name: Acyl carrier protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.64546 KDa |
| Sequence | String: MSTIEERVKK IIGEQLGVKQ EEVTNNASFV EDLGADSLDT VELVMALEEE FDTEIPDEEA EKITTVQAAI DYINGHQA UniProtKB: Acyl carrier protein |
-Macromolecule #3: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 3 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #4: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyl...
| Macromolecule | Name: [(2~{R})-1-[2-azanylethoxy(oxidanyl)phosphoryl]oxy-3-hexadecanoyloxy-propan-2-yl] (~{Z})-octadec-9-enoate type: ligand / ID: 4 / Number of copies: 1 / Formula: 6OU |
|---|---|
| Molecular weight | Theoretical: 717.996 Da |
| Chemical component information |  ChemComp-6OU: |
-Macromolecule #5: 4'-PHOSPHOPANTETHEINE
| Macromolecule | Name: 4'-PHOSPHOPANTETHEINE / type: ligand / ID: 5 / Number of copies: 1 / Formula: PNS |
|---|---|
| Molecular weight | Theoretical: 358.348 Da |
| Chemical component information |  ChemComp-PNS: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 1 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: Solution was filtered and degassed. | |||||||||
| Grid | Model: Homemade / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: OTHER | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 298 K / Instrument: SPOTITON Details: The Spotiton V1.0 robot was operating at room temperature and moderate humidity.. | |||||||||
| Details | Protein was incorporated into lipid nanodiscs. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 15 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Frames/image: 1-90 / Number grids imaged: 4 / Number real images: 7274 / Average exposure time: 13.05 sec. / Average electron dose: 95.72 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6w98: |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)