[English] 日本語
 Yorodumi
Yorodumi- EMDB-21068: Structure of GCP6 in the native human gamma-tubulin ring complex -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21068 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of GCP6 in the native human gamma-tubulin ring complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GCP / GCP6 / gamma-tubulin ring complex / gTuRC / g-TuRC / microtubule / microtubule nucleation / single particle cryo-EM structure / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationgamma-tubulin complex / gamma-tubulin ring complex / microtubule nucleation / gamma-tubulin binding / spindle assembly / cytoplasmic microtubule organization / Recruitment of mitotic centrosome proteins and complexes / Recruitment of NuMA to mitotic centrosomes / meiotic cell cycle / spindle pole ...gamma-tubulin complex / gamma-tubulin ring complex / microtubule nucleation / gamma-tubulin binding / spindle assembly / cytoplasmic microtubule organization / Recruitment of mitotic centrosome proteins and complexes / Recruitment of NuMA to mitotic centrosomes / meiotic cell cycle / spindle pole / mitotic cell cycle / microtubule binding / microtubule / centrosome / membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||
 Authors Authors | Wieczorek M / Urnavicius L | |||||||||
| Funding support |  United States, United States,  France, 2 items France, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: Asymmetric Molecular Architecture of the Human γ-Tubulin Ring Complex. Authors: Michal Wieczorek / Linas Urnavicius / Shih-Chieh Ti / Kelly R Molloy / Brian T Chait / Tarun M Kapoor /  Abstract: The γ-tubulin ring complex (γ-TuRC) is an essential regulator of centrosomal and acentrosomal microtubule formation, yet its structure is not known. Here, we present a cryo-EM reconstruction of the ...The γ-tubulin ring complex (γ-TuRC) is an essential regulator of centrosomal and acentrosomal microtubule formation, yet its structure is not known. Here, we present a cryo-EM reconstruction of the native human γ-TuRC at ∼3.8 Å resolution, revealing an asymmetric, cone-shaped structure. Pseudo-atomic models indicate that GCP4, GCP5, and GCP6 form distinct Y-shaped assemblies that structurally mimic GCP2/GCP3 subcomplexes distal to the γ-TuRC "seam." We also identify an unanticipated structural bridge that includes an actin-like protein and spans the γ-TuRC lumen. Despite its asymmetric architecture, the γ-TuRC arranges γ-tubulins into a helical geometry poised to nucleate microtubules. Diversity in the γ-TuRC subunits introduces large (>100,000 Å) surfaces in the complex that allow for interactions with different regulatory factors. The observed compositional complexity of the γ-TuRC could self-regulate its assembly into a cone-shaped structure to control microtubule formation across diverse contexts, e.g., within biological condensates or alongside existing filaments. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21068.map.gz emd_21068.map.gz | 7.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21068-v30.xml emd-21068-v30.xml emd-21068.xml emd-21068.xml | 14 KB 14 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21068.png emd_21068.png | 41.4 KB | ||
| Filedesc metadata |  emd-21068.cif.gz emd-21068.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21068 http://ftp.pdbj.org/pub/emdb/structures/EMD-21068 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21068 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21068 | HTTPS FTP |
-Related structure data
| Related structure data |  6v6cMC  6v5vC  6v69C  6v6bC  6v6sC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21068.map.gz / Format: CCP4 / Size: 190.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21068.map.gz / Format: CCP4 / Size: 190.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.335 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
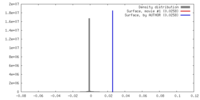
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Native human gamma-tubulin ring complex
| Entire | Name: Native human gamma-tubulin ring complex |
|---|---|
| Components |
|
-Supramolecule #1: Native human gamma-tubulin ring complex
| Supramolecule | Name: Native human gamma-tubulin ring complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Gamma-tubulin complex component 6
| Macromolecule | Name: Gamma-tubulin complex component 6 / type: protein_or_peptide / ID: 1 Details: Please note that the full sequence for GCP6 (Uniprot accession number Q96RT7) was much longer than our modeled region, and the alignment failed to capture the domains we built, despite all ...Details: Please note that the full sequence for GCP6 (Uniprot accession number Q96RT7) was much longer than our modeled region, and the alignment failed to capture the domains we built, despite all residues being numbered accordingly in the submitted coordinates. See below for full sequence. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 200.733641 KDa |
| Sequence | String: MASITQLFDD LCEALLPAAK THLGQRSVNR KRAKRSLKKV AYNALFTNLF QDETQQLQPD MSKLPARNKI LMLSFDLRVG GLGPKADRL EELVEELEAA PCCPLLEVGS VLDLLVQLAG SGPPQVLPRK RDYFLNNKHV GRNVPYSGYD CDDLSVFEMD V QSLISREE ...String: MASITQLFDD LCEALLPAAK THLGQRSVNR KRAKRSLKKV AYNALFTNLF QDETQQLQPD MSKLPARNKI LMLSFDLRVG GLGPKADRL EELVEELEAA PCCPLLEVGS VLDLLVQLAG SGPPQVLPRK RDYFLNNKHV GRNVPYSGYD CDDLSVFEMD V QSLISREE CLCHSMIQET LQVMEAAPGT GLPTVGLFSF GDPCGDRFER DTRVSLFGAL VHSRTYDMDV RLGLPPVPDN AD LSGLAIK VPPSVDQWED EGFQSASNLT PDSQSEPSVT PDVDLWEAAL TYEASKRRCW ERVGCPPGHR EEPYLTEAGR DAF DKFCRL HQGELQLLAG GVLQAPQPVL VKECELVKDV LNVLIGVVSA TFSLCQPAQA FVVKRGVHVS GASPESISSL LSEV AEYGT CYTRLSHFSL QPVLDSLYSK GLVFQAFTSG LRRYLQYYRA CVLSTPPTLS LLTIGFLFKK LGRQLRYLAE LCGVG AVLP GTCGGGPRAA FPTGVKLLSY LYQEALHNCS NEHYPVLLSL LKTSCEPYTR FIHDWVYSGV FRDAYGEFMI QVNHEY LSF RDKLYWTHGY VLISKEVEDC VPVFLKHIAH DIYVCGKTIN LLKLCCPRHY LCWSDVPVPR ISVIFSLEEL KEIEKDC AV YVGRMERVAR HSSVSKEEKE LRMEIAKQEL IAHAREAASR VLSALSDRQM SERMALDARK REQFQRLKEQ FVKDQERR Q AARQEELDDD FSYARELRDR ERRLKSLEEE LERKARQALV DHYSKLSAEA ARREQKALWR IQRHRLESAR LRFLLEDEK HIQEMLKAVS EAHQPQEPPD VLLSVHPQVT SPGPEHPEGG QGCDSGSAEQ HSPAWDGWNR PGLLTPQPLK PLAVGAGGRG LQQAEGARP FSDSLSIGDF LPVGPGAEPS VQTGMVPLLE VALQTINLDL PPSAPGEAPA AASTQPSRPQ EYDFSTVLRP A VATSPAPG PLQAAECSLG SSGLQLWEDS CGKMDACGSA SRETLLPSHP PRRAALEEGS SQPTERLFGQ VSGGGLPTGD YA SEIAPTR PRWNTHGHVS DASIRVGENV SDVAPTQPRW NTHGHVSNAS ISLGESVSDV APTRPRWNIH GHVSNASIRV GEN VSDVAP TRPRWNTHGH VSNASIRVGE NVSDVAPTRP RWNTHGHVSD ASISLGESVS DMAPARPRWN THGHVSDASI SLGE SVSDM APTRPRWNTH GHVSDTSIRV GENVSDVAPI RSRCNTHGHV SDASISLGEP VSDVVSTRPR WNTHVPIPPP HMVLG ALSP EAEPNTPRPQ QSPPGHTSQS ALSLGAQSTV LDCGPRLPVE VGPSLSSPSS GCGEGSISVG ENVSDVAPTQ PWWPNT PGD SVSEELGPGR SGDTEDLSPN WPLNSQEDTA AQSSPGRGEE AEASAAEAQG GEQAYLAGLA GQYHLERYPD SYESMSE PP IAHLLRPVLP RAFAFPVDPQ VQSAADETAV QLSELLTLPV LMKRSITAPL AAHISLVNKA AVDYFFVELH LEAHYEAL R HFLLMEDGEF AQSLSDLLFE KLGAGQTPGE LLNPLVLNSV LSKALQCSLH GDTPHASNLS LALKYLPEVF APNAPDVLS CLELRYKVDW PLNIVITEGC VSKYSGVFSF LLQLKLMMWA LKDVCFHLKR TALLSHMAGS VQFRQLQLFK HEMQHFVKVI QGYIANQIL HVTWCEFRAR LATVGDLEEI QRAHAEYLHK AVFRGLLTEK AAPVMNVIHS IFSLVLKFRS QLISQAWGPP G GPRGAEHP NFALMQQSYN TFKYYSHFLF KVVTKLVNRG YQPHLEDFLL RINFNNYYQD A UniProtKB: Gamma-tubulin complex component 6 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)